A NanoBRET-Based H3R Conformational Biosensor to Study Real-Time H3 Receptor Pharmacology in Cell Membranes and Living Cells
- PMID: 35897787
- PMCID: PMC9332000
- DOI: 10.3390/ijms23158211
A NanoBRET-Based H3R Conformational Biosensor to Study Real-Time H3 Receptor Pharmacology in Cell Membranes and Living Cells
Abstract
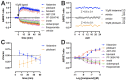
Conformational biosensors to monitor the activation state of G protein-coupled receptors are a useful addition to the molecular pharmacology assay toolbox to characterize ligand efficacy at the level of receptor proteins instead of downstream signaling. We recently reported the initial characterization of a NanoBRET-based conformational histamine H3 receptor (H3R) biosensor that allowed the detection of both (partial) agonism and inverse agonism on living cells in a microplate reader assay format upon stimulation with H3R ligands. In the current study, we have further characterized this H3R biosensor on intact cells by monitoring the effect of consecutive ligand injections in time and evaluating its compatibility with photopharmacological ligands that contain a light-sensitive azobenzene moiety for photo-switching. In addition, we have validated the H3R biosensor in membrane preparations and found that observed potency values better correlated with binding affinity values that were measured in radioligand competition binding assays on membranes. Hence, the H3R conformational biosensor in membranes might be a ready-to-use, high-throughput alternative for radioligand binding assays that in addition can also detect ligand efficacies with comparable values as the intact cell assay.
Keywords: BRET; GPCR; H3R; conformational biosensor; histamine.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


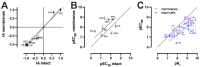
Similar articles
-
Development of a Conformational Histamine H3 Receptor Biosensor for the Synchronous Screening of Agonists and Inverse Agonists.ACS Sens. 2020 Jun 26;5(6):1734-1742. doi: 10.1021/acssensors.0c00397. Epub 2020 May 28. ACS Sens. 2020. PMID: 32397705 Free PMC article.
-
Homogeneous, Real-Time NanoBRET Binding Assays for the Histamine H3 and H4 Receptors on Living Cells.Mol Pharmacol. 2018 Dec;94(6):1371-1381. doi: 10.1124/mol.118.113373. Epub 2018 Sep 24. Mol Pharmacol. 2018. PMID: 30249614
-
Pharmacological characterization of seven human histamine H3 receptor isoforms.Eur J Pharmacol. 2024 Apr 5;968:176450. doi: 10.1016/j.ejphar.2024.176450. Epub 2024 Feb 21. Eur J Pharmacol. 2024. PMID: 38387718
-
Progress in the development of histamine H3 receptor antagonists/inverse agonists: a patent review (2013-2017).Expert Opin Ther Pat. 2018 Mar;28(3):175-196. doi: 10.1080/13543776.2018.1424135. Epub 2018 Jan 15. Expert Opin Ther Pat. 2018. PMID: 29334795 Review.
-
New Horizons on Molecular Pharmacology Applied to Drug Discovery: When Resonance Overcomes Radioligand Binding.Curr Radiopharm. 2017;10(1):16-20. doi: 10.2174/1874471010666170208152420. Curr Radiopharm. 2017. PMID: 28183248 Review.
References
-
- Masureel M., Zou Y., Picard L.-P., Van Der Westhuizen E., Mahoney J.P., Rodrigues J.P.G.L.M., Mildorf T.J., Dror R.O., Shaw D.E., Bouvier M., et al. Structural insights into binding specificity, efficacy and bias of a β2AR partial agonist. Nat. Chem. Biol. 2018;14:1059–1066. doi: 10.1038/s41589-018-0145-x. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

