Automatic Multi-functional Integration Program (AMFIP) towards all-optical mechano-electrophysiology interrogation
- PMID: 35901062
- PMCID: PMC9333221
- DOI: 10.1371/journal.pone.0266098
Automatic Multi-functional Integration Program (AMFIP) towards all-optical mechano-electrophysiology interrogation
Abstract
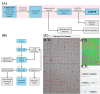
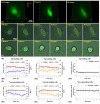
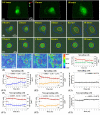
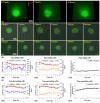
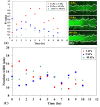
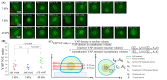
Automatic operations of multi-functional and time-lapse live-cell imaging are necessary for the biomedical science community to study active, multi-faceted, and long-term biological phenomena. To achieve automatic control, most existing solutions often require the purchase of extra software programs and hardware that rely on the manufacturers' own specifications. However, these software programs are usually non-user-programmable and unaffordable for many laboratories. To address this unmet need, we have developed a novel open-source software program, titled Automatic Multi-functional Integration Program (AMFIP), as a new Java-based and hardware-independent system that provides proven advantages over existing alternatives to the scientific community. Without extra hardware, AMFIP enables the functional synchronization of the μManager software platform, the Nikon NIS-Elements platform, and other 3rd party software to achieve automatic operations of most commercially available microscopy systems, including but not limited to those from Nikon. AMFIP provides a user-friendly and programmable graphical user interface (GUI), opening the door to expanding the customizability for myriad hardware and software systems according to user-specific experimental requirements and environments. To validate the intended purposes of developing AMFIP, we applied it to elucidate the question whether single cells, prior to their full spreading, can sense and respond to a soft solid substrate, and if so, how does the interaction depend on the cell spreading time and the stiffness of the substrate. Using a CRISPR/Cas9-engineered human epithelial Beas2B (B2B) cell line that expresses mNeonGreen2-tagged mechanosensitive Yes-associated protein (YAP), we show that single B2B cells develop distinct substrate-stiffness-dependent YAP expressions within 10 hours at most on the substrate, suggesting that cells are able to sense, distinguish, and respond to mechanical cues prior to the establishment of full cell spreading. In summary, AMFIP provides a reliable, open-source, and cost-free solution that has the validated long-term utility to satisfy the need of automatic imaging operations in the scientific community.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
All-optical Mechanobiology Interrogation of Yes-associated Protein in Human Cancer and Normal Cells using a Multi-functional System.J Vis Exp. 2021 Dec 20;(178). doi: 10.3791/62934. J Vis Exp. 2021. PMID: 34978284
-
Advanced methods of microscope control using μManager software.J Biol Methods. 2014;1(2):e10. doi: 10.14440/jbm.2014.36. J Biol Methods. 2014. PMID: 25606571 Free PMC article.
-
An Integrated Platform for In Vivo Electrophysiology in Spatial Cognition Experiments.eNeuro. 2023 Nov 21;10(11):ENEURO.0274-23.2023. doi: 10.1523/ENEURO.0274-23.2023. Print 2023 Nov. eNeuro. 2023. PMID: 37989581 Free PMC article.
-
Applications and challenges of digital pathology and whole slide imaging.Biotech Histochem. 2015 Jul;90(5):341-7. doi: 10.3109/10520295.2015.1044566. Epub 2015 May 15. Biotech Histochem. 2015. PMID: 25978139 Review.
-
MPScope 2.0: A Computer System for Two-Photon Laser Scanning Microscopy with Concurrent Plasma-Mediated Ablation and Electrophysiology.In: Frostig RD, editor. In Vivo Optical Imaging of Brain Function. 2nd edition. Boca Raton (FL): CRC Press/Taylor & Francis; 2009. Chapter 4. In: Frostig RD, editor. In Vivo Optical Imaging of Brain Function. 2nd edition. Boca Raton (FL): CRC Press/Taylor & Francis; 2009. Chapter 4. PMID: 26844327 Free Books & Documents. Review.
Cited by
-
YAP at the Crossroads of Biomechanics and Drug Resistance in Human Cancer.Int J Mol Sci. 2023 Aug 6;24(15):12491. doi: 10.3390/ijms241512491. Int J Mol Sci. 2023. PMID: 37569866 Free PMC article. Review.
References
-
- Tang X, Bajaj P, Bashir R, Saif T. How far cardiac cells can see each other mechanically. Soft Matter. 2011;7(13):6151–8.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

