Skeletal progenitors preserve proliferation and self-renewal upon inhibition of mitochondrial respiration by rerouting the TCA cycle
- PMID: 35905715
- PMCID: PMC9380255
- DOI: 10.1016/j.celrep.2022.111105
Skeletal progenitors preserve proliferation and self-renewal upon inhibition of mitochondrial respiration by rerouting the TCA cycle
Abstract
A functional electron transport chain (ETC) is crucial for supporting bioenergetics and biosynthesis. Accordingly, ETC inhibition decreases proliferation in cancer cells but does not seem to impair stem cell proliferation. However, it remains unclear how stem cells metabolically adapt. In this study, we show that pharmacological inhibition of complex III of the ETC in skeletal stem and progenitor cells induces glycolysis side pathways and reroutes the tricarboxylic acid (TCA) cycle to regenerate NAD+ and preserve cell proliferation. These metabolic changes also culminate in increased succinate and 2-hydroxyglutarate levels that inhibit Ten-eleven translocation (TET) DNA demethylase activity, thereby preserving self-renewal and multilineage potential. Mechanistically, mitochondrial malate dehydrogenase and reverse succinate dehydrogenase activity proved to be essential for the metabolic rewiring in response to ETC inhibition. Together, these data show that the metabolic plasticity of skeletal stem and progenitor cells allows them to bypass ETC blockade and preserve their self-renewal.
Keywords: CP: Metabolism; CP: Stem cell research; NAD regeneration; TCA rerouting; TET activity; cell-based regenerative medicine; electron transport chain; metabolic plasticity; proliferation; reverse succinate dehydrogenase; self-renewal; skeletal stem cells.
Copyright © 2022 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


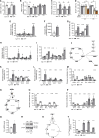





Similar articles
-
Carnosine Inhibits the Proliferation of Human Cervical Gland Carcinoma Cells Through Inhibiting Both Mitochondrial Bioenergetics and Glycolysis Pathways and Retarding Cell Cycle Progression.Integr Cancer Ther. 2018 Mar;17(1):80-91. doi: 10.1177/1534735416684551. Epub 2016 Dec 23. Integr Cancer Ther. 2018. PMID: 28008780 Free PMC article.
-
Reversal of mitochondrial malate dehydrogenase 2 enables anaplerosis via redox rescue in respiration-deficient cells.Mol Cell. 2022 Dec 1;82(23):4537-4547.e7. doi: 10.1016/j.molcel.2022.10.005. Epub 2022 Nov 2. Mol Cell. 2022. PMID: 36327975
-
GLUD1 determines murine muscle stem cell fate by controlling mitochondrial glutamate levels.Dev Cell. 2024 Nov 4;59(21):2850-2865.e8. doi: 10.1016/j.devcel.2024.07.015. Epub 2024 Aug 8. Dev Cell. 2024. PMID: 39121856
-
Mitochondrial dysfunctions in cancer: genetic defects and oncogenic signaling impinging on TCA cycle activity.Cancer Lett. 2015 Jan 28;356(2 Pt A):217-23. doi: 10.1016/j.canlet.2014.02.023. Epub 2014 Mar 12. Cancer Lett. 2015. PMID: 24614286 Review.
-
Respiratory metabolism: glycolysis, the TCA cycle and mitochondrial electron transport.Curr Opin Plant Biol. 2004 Jun;7(3):254-61. doi: 10.1016/j.pbi.2004.03.007. Curr Opin Plant Biol. 2004. PMID: 15134745 Review.
Cited by
-
Metabolic reprogramming in skeletal cell differentiation.Bone Res. 2024 Oct 11;12(1):57. doi: 10.1038/s41413-024-00374-0. Bone Res. 2024. PMID: 39394187 Free PMC article. Review.
-
Oxidative phosphorylation in bone cells.Bone Rep. 2023 May 23;18:101688. doi: 10.1016/j.bonr.2023.101688. eCollection 2023 Jun. Bone Rep. 2023. PMID: 37275785 Free PMC article.
-
Stirred culture of cartilaginous microtissues promotes chondrogenic hypertrophy through exposure to intermittent shear stress.Bioeng Transl Med. 2022 Dec 29;8(3):e10468. doi: 10.1002/btm2.10468. eCollection 2023 May. Bioeng Transl Med. 2022. PMID: 37206246 Free PMC article.
-
Proteomic Analysis of the Mitochondrial Responses in P19 Embryonic Stem Cells Exposed to Florfenicol.Toxics. 2023 Dec 6;11(12):992. doi: 10.3390/toxics11120992. Toxics. 2023. PMID: 38133393 Free PMC article.
-
HIF1 activation safeguards cortical bone formation against impaired oxidative phosphorylation.JCI Insight. 2024 Aug 1;9(18):e182330. doi: 10.1172/jci.insight.182330. JCI Insight. 2024. PMID: 39088272 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

