Transcription factor network analysis identifies REST/NRSF as an intrinsic regulator of CNS regeneration in mice
- PMID: 35906210
- PMCID: PMC9338053
- DOI: 10.1038/s41467-022-31960-7
Transcription factor network analysis identifies REST/NRSF as an intrinsic regulator of CNS regeneration in mice
Abstract
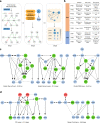
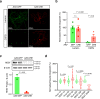
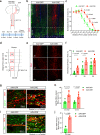
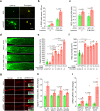
The inability of neurons to regenerate long axons within the CNS is a major impediment to improving outcome after spinal cord injury, stroke, and other CNS insults. Recent advances have uncovered an intrinsic program that involves coordinate regulation by multiple transcription factors that can be manipulated to enhance growth in the peripheral nervous system. Here, we use a systems genomics approach to characterize regulatory relationships of regeneration-associated transcription factors, identifying RE1-Silencing Transcription Factor (REST; Neuron-Restrictive Silencer Factor, NRSF) as a predicted upstream suppressor of a pro-regenerative gene program associated with axon regeneration in the CNS. We validate our predictions using multiple paradigms, showing that mature mice bearing cell type-specific deletions of REST or expressing dominant-negative mutant REST show improved regeneration of the corticospinal tract and optic nerve after spinal cord injury and optic nerve crush, which is accompanied by upregulation of regeneration-associated genes in cortical motor neurons and retinal ganglion cells, respectively. These analyses identify a role for REST as an upstream suppressor of the intrinsic regenerative program in the CNS and demonstrate the utility of a systems biology approach involving integrative genomics and bio-informatics to prioritize hypotheses relevant to CNS repair.
© 2022. The Author(s).
Conflict of interest statement
Z.H. is a co-founder of Rugen and Myro Therapeutics. The remaining authors declare no competing interests.
Figures




References
-
- Cajal, S. R. Cajal’s degeneration and regeneration of the nervous system. (History of Neuroscience, 1991).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

