Single-cell transcriptomic analysis of vascular endothelial cells in zebrafish embryos
- PMID: 35906287
- PMCID: PMC9338088
- DOI: 10.1038/s41598-022-17127-w
Single-cell transcriptomic analysis of vascular endothelial cells in zebrafish embryos
Abstract
Vascular endothelial cells exhibit substantial phenotypic and transcriptional heterogeneity which is established during early embryogenesis. However, the molecular mechanisms involved in establishing endothelial cell diversity are still not well understood. Zebrafish has emerged as an advantageous model to study vascular development. Despite its importance, the single-cell transcriptomic profile of vascular endothelial cells during zebrafish development is still missing. To address this, we applied single-cell RNA-sequencing (scRNA-seq) of vascular endothelial cells isolated from zebrafish embryos at the 24 hpf stage. Six distinct clusters or subclusters related to vascular endothelial cells were identified which include arterial, two venous, cranial, endocardial and endothelial progenitor cell subtypes. Furthermore, we validated our findings by characterizing novel markers for arterial, venous, and endocardial cells. We experimentally confirmed the presence of two transcriptionally different venous cell subtypes, demonstrating heterogeneity among venous endothelial cells at this early developmental stage. This dataset will be a valuable resource for future functional characterization of vascular endothelial cells and interrogation of molecular mechanisms involved in the establishment of their heterogeneity and cell-fate decisions.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

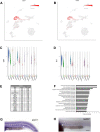
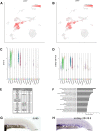
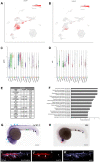

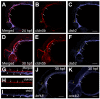
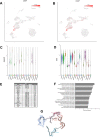

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

