Highly Overexpressed AtC3H18 Impairs Microgametogenesis via Promoting the Continuous Assembly of mRNP Granules
- PMID: 35909782
- PMCID: PMC9335048
- DOI: 10.3389/fpls.2022.932793
Highly Overexpressed AtC3H18 Impairs Microgametogenesis via Promoting the Continuous Assembly of mRNP Granules
Abstract
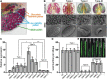
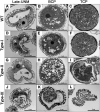
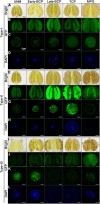
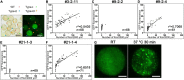
Plant CCCH zinc-finger proteins form a large family of regulatory proteins function in many aspects of plant growth, development and environmental responses. Despite increasing reports indicate that many CCCH zinc-finger proteins exhibit similar subcellular localization of being localized in cytoplasmic foci, the underlying molecular mechanism and the connection between this specific localization pattern and protein functions remain largely elusive. Here, we identified another cytoplasmic foci-localized CCCH zinc-finger protein, AtC3H18, in Arabidopsis thaliana. AtC3H18 is predominantly expressed in developing pollen during microgametogenesis. Although atc3h18 mutants did not show any abnormal phenotype, possibly due to redundant gene(s), aberrant AtC3H18 expression levels caused by overexpression resulted in the assembly of AtC3H18-positive granules in a dose-dependent manner, which in turn led to male sterility phenotype, highlighting the importance of fine-tuned AtC3H18 expression. Further analyzes demonstrated that AtC3H18-positive granules are messenger ribonucleoprotein (mRNP) granules, since they can exhibit liquid-like physical properties, and are associated with another two mRNP granules known as processing bodies (PBs) and stress granules (SGs), reservoirs of translationally inhibited mRNAs. Moreover, the assembly of AtC3H18-positive granules depends on mRNA availability. Combined with our previous findings on the AtC3H18 homologous genes in Brassica campestris, we concluded that appropriate expression level of AtC3H18 during microgametogenesis is essential for normal pollen development, and we also speculated that AtC3H18 may act as a key component of mRNP granules to modulate pollen mRNAs by regulating the assembly/disassembly of mRNP granules, thereby affecting pollen development.
Keywords: Arabidopsis thaliana; AtC3H18; mRNP granules; pollen development; processing bodies (PBs); stress granules (SGs).
Copyright © 2022 Xu, Liu, Xiong, Shen, Huang, Yu and Cao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The handling editor declared a past collaboration with one of the authors, LH.
Figures




Similar articles
-
A LlMYB305-LlC3H18-LlWRKY33 module regulates thermotolerance in lily.Mol Hortic. 2023 Aug 17;3(1):15. doi: 10.1186/s43897-023-00064-1. Mol Hortic. 2023. PMID: 37789438 Free PMC article.
-
Heterologous Expression of Two Brassica campestris CCCH Zinc-Finger Proteins in Arabidopsis Induces Cytoplasmic Foci and Causes Pollen Abortion.Int J Mol Sci. 2023 Nov 28;24(23):16862. doi: 10.3390/ijms242316862. Int J Mol Sci. 2023. PMID: 38069184 Free PMC article.
-
AtC3H18L is a stop-codon read-through gene and encodes a novel non-tandem CCCH zinc-finger protein that can form cytoplasmic foci similar to mRNP granules.Biochem Biophys Res Commun. 2020 Jul 12;528(1):140-145. doi: 10.1016/j.bbrc.2020.05.081. Epub 2020 May 23. Biochem Biophys Res Commun. 2020. PMID: 32451083
-
mRNP granules. Assembly, function, and connections with disease.RNA Biol. 2014;11(8):1019-30. doi: 10.4161/15476286.2014.972208. RNA Biol. 2014. PMID: 25531407 Free PMC article. Review.
-
Stress Granules and Processing Bodies in Translational Control.Cold Spring Harb Perspect Biol. 2019 May 1;11(5):a032813. doi: 10.1101/cshperspect.a032813. Cold Spring Harb Perspect Biol. 2019. PMID: 30082464 Free PMC article. Review.
Cited by
-
A LlMYB305-LlC3H18-LlWRKY33 module regulates thermotolerance in lily.Mol Hortic. 2023 Aug 17;3(1):15. doi: 10.1186/s43897-023-00064-1. Mol Hortic. 2023. PMID: 37789438 Free PMC article.
-
Transcription Factors and Their Regulatory Roles in the Male Gametophyte Development of Flowering Plants.Int J Mol Sci. 2024 Jan 1;25(1):566. doi: 10.3390/ijms25010566. Int J Mol Sci. 2024. PMID: 38203741 Free PMC article. Review.
-
Heterologous Expression of Two Brassica campestris CCCH Zinc-Finger Proteins in Arabidopsis Induces Cytoplasmic Foci and Causes Pollen Abortion.Int J Mol Sci. 2023 Nov 28;24(23):16862. doi: 10.3390/ijms242316862. Int J Mol Sci. 2023. PMID: 38069184 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

