Targeting SREBP-1-Mediated Lipogenesis as Potential Strategies for Cancer
- PMID: 35912181
- PMCID: PMC9330218
- DOI: 10.3389/fonc.2022.952371
Targeting SREBP-1-Mediated Lipogenesis as Potential Strategies for Cancer
Abstract
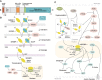
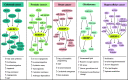
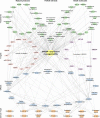
Sterol regulatory element binding protein-1 (SREBP-1), a transcription factor with a basic helix-loop-helix leucine zipper, has two isoforms, SREBP-1a and SREBP-1c, derived from the same gene for regulating the genes of lipogenesis, including acetyl-CoA carboxylase, fatty acid synthase, and stearoyl-CoA desaturase. Importantly, SREBP-1 participates in metabolic reprogramming of various cancers and has been a biomarker for the prognosis or drug efficacy for the patients with cancer. In this review, we first introduced the structure, activation, and key upstream signaling pathway of SREBP-1. Then, the potential targets and molecular mechanisms of SREBP-1-regulated lipogenesis in various types of cancer, such as colorectal, prostate, breast, and hepatocellular cancer, were summarized. We also discussed potential therapies targeting the SREBP-1-regulated pathway by small molecules, natural products, or the extracts of herbs against tumor progression. This review could provide new insights in understanding advanced findings about SREBP-1-mediated lipogenesis in cancer and its potential as a target for cancer therapeutics.
Keywords: SREBP-1; cancer therapy; fatty acid synthase; fatty acids; lipogenesis.
Copyright © 2022 Zhao, Lin and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Targeting SREBP-2-Regulated Mevalonate Metabolism for Cancer Therapy.Front Oncol. 2020 Aug 21;10:1510. doi: 10.3389/fonc.2020.01510. eCollection 2020. Front Oncol. 2020. PMID: 32974183 Free PMC article. Review.
-
Novel insights for SREBP-1 as a key transcription factor in regulating lipogenesis in a freshwater teleost, grass carp Ctenopharyngodon idella.Br J Nutr. 2019 Dec 14;122(11):1201-1211. doi: 10.1017/S0007114519001934. Br J Nutr. 2019. PMID: 31782376
-
Inhibition of lipogenesis and induction of apoptosis by valproic acid in prostate cancer cells via the C/EBPα/SREBP-1 pathway.Acta Biochim Biophys Sin (Shanghai). 2021 Mar 2;53(3):354-364. doi: 10.1093/abbs/gmab002. Acta Biochim Biophys Sin (Shanghai). 2021. PMID: 33471067
-
Exendin-4 Inhibits Hepatic Lipogenesis by Increasing β-Catenin Signaling.PLoS One. 2016 Dec 1;11(12):e0166913. doi: 10.1371/journal.pone.0166913. eCollection 2016. PLoS One. 2016. PMID: 27907035 Free PMC article.
-
Hepatic steatosis: a role for de novo lipogenesis and the transcription factor SREBP-1c.Diabetes Obes Metab. 2010 Oct;12 Suppl 2:83-92. doi: 10.1111/j.1463-1326.2010.01275.x. Diabetes Obes Metab. 2010. PMID: 21029304 Review.
Cited by
-
IRE1α-XBP1s axis regulates SREBP1-dependent MRP1 expression to promote chemoresistance in non-small cell lung cancer cells.Thorac Cancer. 2024 Oct;15(29):2116-2127. doi: 10.1111/1759-7714.15442. Epub 2024 Sep 8. Thorac Cancer. 2024. PMID: 39245881 Free PMC article.
-
Roles of posttranslational modifications in lipid metabolism and cancer progression.Biomark Res. 2024 Nov 18;12(1):141. doi: 10.1186/s40364-024-00681-y. Biomark Res. 2024. PMID: 39551780 Free PMC article. Review.
-
Enhancing doxorubicin's anticancer impact in colorectal cancer by targeting the Akt/Gsk3β/mTOR-SREBP1 signaling axis with an HDAC inhibitor.Korean J Physiol Pharmacol. 2025 May 1;29(3):321-335. doi: 10.4196/kjpp.24.274. Korean J Physiol Pharmacol. 2025. PMID: 40254556 Free PMC article.
-
PTHrP Regulates Fatty Acid Metabolism via Novel lncRNA in Breast Cancer Initiation and Progression Models.Cancers (Basel). 2023 Jul 25;15(15):3763. doi: 10.3390/cancers15153763. Cancers (Basel). 2023. PMID: 37568579 Free PMC article.
-
The cooperative regulatory effect of the miRNA-130 family on milk fat metabolism in dairy cows.Anim Biosci. 2024 Jul;37(7):1289-1302. doi: 10.5713/ab.23.0485. Epub 2024 Apr 25. Anim Biosci. 2024. PMID: 38665085 Free PMC article.
References
-
- Hua X, Yokoyama C, Wu J, Briggs MR, Brown MS, Goldstein JL, et al. . Srebp-2, a Second Basic-Helix-Loop-Helix-Leucine Zipper Protein That Stimulates Transcription by Binding to a Sterol Regulatory Element. Proc Natl Acad Sci USA (1993) 90(24):11603–7. doi: 10.1073/pnas.90.24.11603 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

