Chemical interference with DSIF complex formation lowers synthesis of mutant huntingtin gene products and curtails mutant phenotypes
- PMID: 35914128
- PMCID: PMC9371670
- DOI: 10.1073/pnas.2204779119
Chemical interference with DSIF complex formation lowers synthesis of mutant huntingtin gene products and curtails mutant phenotypes
Abstract
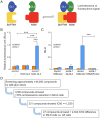
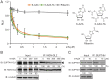
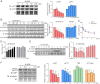
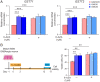
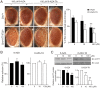
Earlier work has shown that siRNA-mediated reduction of the SUPT4H or SUPT5H proteins, which interact to form the DSIF complex and facilitate transcript elongation by RNA polymerase II (RNAPII), can decrease expression of mutant gene alleles containing nucleotide repeat expansions differentially. Using luminescence and fluorescence assays, we identified chemical compounds that interfere with the SUPT4H-SUPT5H interaction and then investigated their effects on synthesis of mRNA and protein encoded by mutant alleles containing repeat expansions in the huntingtin gene (HTT), which causes the inherited neurodegenerative disorder, Huntington's Disease (HD). Here we report that such chemical interference can differentially affect expression of HTT mutant alleles, and that a prototypical chemical, 6-azauridine (6-AZA), that targets the SUPT4H-SUPT5H interaction can modify the biological response to mutant HTT gene expression. Selective and dose-dependent effects of 6-AZA on expression of HTT alleles containing nucleotide repeat expansions were seen in multiple types of cells cultured in vitro, and in a Drosophila melanogaster animal model for HD. Lowering of mutant HD protein and mitigation of the Drosophila "rough eye" phenotype associated with degeneration of photoreceptor neurons in vivo were observed. Our findings indicate that chemical interference with DSIF complex formation can decrease biochemical and phenotypic effects of nucleotide repeat expansions.
Keywords: DSIF; Huntington’s disease; SUPT4H; Spt4; nucleotide repeats.
Conflict of interest statement
The authors declare no competing interest.
Figures
Similar articles
-
Effects on murine behavior and lifespan of selectively decreasing expression of mutant huntingtin allele by supt4h knockdown.PLoS Genet. 2015 Mar 11;11(3):e1005043. doi: 10.1371/journal.pgen.1005043. eCollection 2015 Mar. PLoS Genet. 2015. PMID: 25760041 Free PMC article.
-
DSIF modulates RNA polymerase II occupancy according to template G + C content.NAR Genom Bioinform. 2022 Jul 27;4(3):lqac054. doi: 10.1093/nargab/lqac054. eCollection 2022 Sep. NAR Genom Bioinform. 2022. PMID: 35910045 Free PMC article.
-
Allele-selective lowering of mutant HTT protein by HTT-LC3 linker compounds.Nature. 2019 Nov;575(7781):203-209. doi: 10.1038/s41586-019-1722-1. Epub 2019 Oct 30. Nature. 2019. PMID: 31666698
-
Huntingtin Lowering Strategies.Int J Mol Sci. 2020 Mar 20;21(6):2146. doi: 10.3390/ijms21062146. Int J Mol Sci. 2020. PMID: 32245050 Free PMC article. Review.
-
Cancer: From Wild-Type to Mutant Huntingtin.J Huntingtons Dis. 2018;7(3):201-208. doi: 10.3233/JHD-180290. J Huntingtons Dis. 2018. PMID: 29889077 Free PMC article. Review.
Cited by
-
Lowering mutant huntingtin by small molecules relieves Huntington's disease symptoms and progression.EMBO Mol Med. 2024 Mar;16(3):523-546. doi: 10.1038/s44321-023-00020-y. Epub 2024 Feb 19. EMBO Mol Med. 2024. PMID: 38374466 Free PMC article.
-
The organophotocatalytic trifluoromethylation of 6-azauracils.RSC Adv. 2025 Apr 10;15(15):11370-11376. doi: 10.1039/d5ra00743g. eCollection 2025 Apr 9. RSC Adv. 2025. PMID: 40213620 Free PMC article.
-
Mutant-Huntingtin Molecular Pathways Elucidate New Targets for Drug Repurposing.Int J Mol Sci. 2023 Nov 27;24(23):16798. doi: 10.3390/ijms242316798. Int J Mol Sci. 2023. PMID: 38069121 Free PMC article. Review.
-
Decoding Nucleotide Repeat Expansion Diseases: Novel Insights from Drosophila melanogaster Studies.Int J Mol Sci. 2024 Nov 2;25(21):11794. doi: 10.3390/ijms252111794. Int J Mol Sci. 2024. PMID: 39519345 Free PMC article. Review.
-
The transcription elongation factors Spt4 and Spt5 control neural progenitor proliferation and are implicated in neuronal remodeling during Drosophila mushroom body development.Front Cell Dev Biol. 2024 Oct 9;12:1434168. doi: 10.3389/fcell.2024.1434168. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39445331 Free PMC article.
References
-
- Yamaguchi Y., et al. , Structure and function of the human transcription elongation factor DSIF. J. Biol. Chem. 274, 8085–8092 (1999). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

