An analysis of the significance of the Tre2/Bub2/CDC 16 (TBC) domain protein family 8 in colorectal cancer
- PMID: 35918393
- PMCID: PMC9345998
- DOI: 10.1038/s41598-022-15629-1
An analysis of the significance of the Tre2/Bub2/CDC 16 (TBC) domain protein family 8 in colorectal cancer
Abstract
The TBC (Tre-2/Bub2/Cdc16, TBC) structural domain is now considered as one of the factors potentially regulating tumor progression. However, to date, studies on the relationship between TBC structural domains and tumors are limited. In this study, we identified the role of TBC1 domain family member 8 (TBC1D8) as an oncogene in colorectal cancer (CRC) by least absolute shrinkage and selection operator (LASSO) and Cox regression analysis, showing that TBC1D8 may independently predict CRC outcome. Functional enrichment and single-cell analysis showed that TBC1D8 levels were associated with hypoxia. TBC1D8 levels were also positively correlated with M2 macrophage infiltration, which may have a complex association with hypoxia. Taken together, these results show that the TBC1D8 gene is involved in colorectal carcinogenesis, and the underlying molecular mechanisms may include hypoxia and immune cell infiltration.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





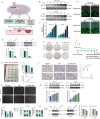


Similar articles
-
Screening for target Rabs of TBC (Tre-2/Bub2/Cdc16) domain-containing proteins based on their Rab-binding activity.Genes Cells. 2006 Sep;11(9):1023-37. doi: 10.1111/j.1365-2443.2006.00997.x. Genes Cells. 2006. PMID: 16923123
-
TBC proteins: GAPs for mammalian small GTPase Rab?Biosci Rep. 2011 Jun;31(3):159-68. doi: 10.1042/BSR20100112. Biosci Rep. 2011. PMID: 21250943 Review.
-
TBC-domain GAPs for Rab GTPases accelerate GTP hydrolysis by a dual-finger mechanism.Nature. 2006 Jul 20;442(7100):303-6. doi: 10.1038/nature04847. Nature. 2006. PMID: 16855591
-
TBC domain family, member 15 is a novel mammalian Rab GTPase-activating protein with substrate preference for Rab7.Biochem Biophys Res Commun. 2005 Sep 16;335(1):154-61. doi: 10.1016/j.bbrc.2005.07.070. Biochem Biophys Res Commun. 2005. PMID: 16055087
-
The critical roles of TBC proteins in human diseases.Yi Chuan. 2018 Jan 20;40(1):12-21. doi: 10.16288/j.yczz.17-343. Yi Chuan. 2018. PMID: 29367189 Review.
Cited by
-
ZSCAN16 expedites hepatocellular carcinoma progression via activating TBC1D31.Cell Div. 2024 Nov 7;19(1):31. doi: 10.1186/s13008-024-00135-9. Cell Div. 2024. PMID: 39511655 Free PMC article.
-
NO enhances the adaptability to high-salt environments by regulating osmotic balance, antioxidant defense, and ion homeostasis in eelgrass based on transcriptome and metabolome analysis.Front Plant Sci. 2024 Feb 7;15:1343154. doi: 10.3389/fpls.2024.1343154. eCollection 2024. Front Plant Sci. 2024. PMID: 38384762 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

