Genomic evidence for global ocean plankton biogeography shaped by large-scale current systems
- PMID: 35920817
- PMCID: PMC9348854
- DOI: 10.7554/eLife.78129
Genomic evidence for global ocean plankton biogeography shaped by large-scale current systems
Abstract
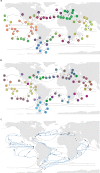
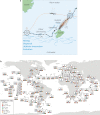
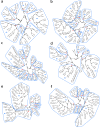
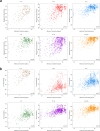
Biogeographical studies have traditionally focused on readily visible organisms, but recent technological advances are enabling analyses of the large-scale distribution of microscopic organisms, whose biogeographical patterns have long been debated. Here we assessed the global structure of plankton geography and its relation to the biological, chemical, and physical context of the ocean (the 'seascape') by analyzing metagenomes of plankton communities sampled across oceans during the Tara Oceans expedition, in light of environmental data and ocean current transport. Using a consistent approach across organismal sizes that provides unprecedented resolution to measure changes in genomic composition between communities, we report a pan-ocean, size-dependent plankton biogeography overlying regional heterogeneity. We found robust evidence for a basin-scale impact of transport by ocean currents on plankton biogeography, and on a characteristic timescale of community dynamics going beyond simple seasonality or life history transitions of plankton.
Keywords: ecology; genetics; genomics; metabarcoding; metagenomics; microbial oceanography; plankton biogeography.
Plain language summary
Oceans are brimming with life invisible to our eyes, a myriad of species of bacteria, viruses and other microscopic organisms essential for the health of the planet. These ‘marine plankton’ are unable to swim against currents and should therefore be constantly on the move, yet previous studies have suggested that distinct species of plankton may in fact inhabit different oceanic regions. However, proving this theory has been challenging; collecting plankton is logistically difficult, and it is often impossible to distinguish between species simply by examining them under a microscope. However, within the last decade, a research schooner called Tara has travelled the globe to gather thousands of plankton samples. At the same time, advances in genomics have made it possible to identify species based only on fragments of their DNA sequence. To understand the hidden geography of plankton communities in Earth’s oceans, Richter et al. pored over DNA from the Tara Oceans expedition. This revealed that, despite being unable to resist the flow of water, various planktonic species which live close to the surface manage to occupy distinct, stable provinces shaped by currents. Different sizes of plankton are distributed in different sized provinces, with the smallest organisms tending to inhabit the smallest areas. Comparing DNA similarities and speeds of currents at the ocean surface revealed how these might stretch and mix plankton communities. Plankton play a critical role in the health of the ocean and the chemical cycles of planet Earth. These results could allow deeper investigation by marine modellers, ecologists, and evolutionary biologists. Meanwhile, work is already underway to investigate how climate change might impact this hidden geography.
© 2022, Richter, Watteaux, Vannier et al.
Conflict of interest statement
DR, RW, TV, JL, PF, GR, NM, NH, GB, OD, TD, AF, SS, RN, CB, DE, FG, LG, KL, EM, JP, SR, SR, CD, SK, MP, SS, SP, JA, JB, CL, EP, PB, SS, FL, LK, CB, MS, EK, MM, IP, PP, PW, Cd, MR, DI, OJ No competing interests declared
Figures







Similar articles
-
Plankton biogeography in the 21st century and impacts of climate change: advances through genomics.C R Biol. 2023 Feb 2;346:13-24. doi: 10.5802/crbiol.107. C R Biol. 2023. PMID: 37254792 Review.
-
Ubiquitous abundance distribution of non-dominant plankton across the global ocean.Nat Ecol Evol. 2018 Aug;2(8):1243-1249. doi: 10.1038/s41559-018-0587-2. Epub 2018 Jun 18. Nat Ecol Evol. 2018. PMID: 29915345
-
Viral to metazoan marine plankton nucleotide sequences from the Tara Oceans expedition.Sci Data. 2017 Aug 1;4:170093. doi: 10.1038/sdata.2017.93. Sci Data. 2017. PMID: 28763055 Free PMC article.
-
Ocean plankton. Patterns and ecological drivers of ocean viral communities.Science. 2015 May 22;348(6237):1261498. doi: 10.1126/science.1261498. Science. 2015. PMID: 25999515
-
Tara Oceans: towards global ocean ecosystems biology.Nat Rev Microbiol. 2020 Aug;18(8):428-445. doi: 10.1038/s41579-020-0364-5. Epub 2020 May 12. Nat Rev Microbiol. 2020. PMID: 32398798 Review.
Cited by
-
Ocean-wide comparisons of mesopelagic planktonic community structures.ISME Commun. 2023 Aug 18;3(1):83. doi: 10.1038/s43705-023-00279-9. ISME Commun. 2023. PMID: 37596349 Free PMC article.
-
Biogeographical and Biodiversity Patterns of Marine Planktonic Bacteria Spanning from the South China Sea across the Gulf of Bengal to the Northern Arabian Sea.Microbiol Spectr. 2023 Jun 15;11(3):e0039823. doi: 10.1128/spectrum.00398-23. Epub 2023 Apr 26. Microbiol Spectr. 2023. PMID: 37098981 Free PMC article.
-
An INDEL genomic approach to explore population diversity of phytoplankton.BMC Genomics. 2024 Nov 6;25(1):1045. doi: 10.1186/s12864-024-10896-w. BMC Genomics. 2024. PMID: 39506649 Free PMC article.
-
Global biogeography of the smallest plankton across ocean depths.Sci Adv. 2023 Nov 10;9(45):eadg9763. doi: 10.1126/sciadv.adg9763. Epub 2023 Nov 8. Sci Adv. 2023. PMID: 37939185 Free PMC article.
-
Scratching the ocean surface: Researchers want to better understand the nature and dynamics of the abundant life living on and in the ocean's surface layers.EMBO Rep. 2023 Sep 6;24(9):e57928. doi: 10.15252/embr.202357928. Epub 2023 Aug 17. EMBO Rep. 2023. PMID: 37589208 Free PMC article.
References
-
- Alberti A, Poulain J, Engelen S, Labadie K, Romac S, Ferrera I, Albini G, Aury JM, Belser C, Bertrand A, Cruaud C, Da Silva C, Dossat C, Gavory F, Gas S, Guy J, Haquelle M, Jacoby E, Jaillon O, Lemainque A, Pelletier E, Samson G, Wessner M, Acinas SG, Royo-Llonch M, Cornejo-Castillo FM, Logares R, Fernández-Gómez B, Bowler C, Cochrane G, Amid C, Hoopen PT, De Vargas C, Grimsley N, Desgranges E, Kandels-Lewis S, Ogata H, Poulton N, Sieracki ME, Stepanauskas R, Sullivan MB, Brum JR, Duhaime MB, Poulos BT, Hurwitz BL, Pesant S, Karsenti E, Wincker P, Genoscope Technical Team. Tara Oceans Consortium Coordinators Viral to metazoan marine plankton nucleotide sequences from the Tara Oceans expedition. Scientific Data. 2017;4:170093. doi: 10.1038/sdata.2017.93. - DOI - PMC - PubMed
-
- Baker FB, Hubert LJ. Measuring the power of hierarchical cluster analysis. Journal of the American Statistical Association. 1975;70:31–38. doi: 10.1080/01621459.1975.10480256. - DOI
-
- Becker RA, Wilks AR, Brownrigg R, Minka TP, Deckmyn A. Draw Geographical Maps. 3.4.0R Package Maps. 2018 https://cran.r-project.org/web/packages/maps/maps.pdf
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources

