Rapamycin rescues mitochondrial dysfunction in cells carrying the m.8344A > G mutation in the mitochondrial tRNALys
- PMID: 35922766
- PMCID: PMC9347137
- DOI: 10.1186/s10020-022-00519-z
Rapamycin rescues mitochondrial dysfunction in cells carrying the m.8344A > G mutation in the mitochondrial tRNALys
Abstract
Background: Myoclonus, Epilepsy and Ragged-Red-Fibers (MERRF) is a mitochondrial encephalomyopathy due to heteroplasmic mutations in mitochondrial DNA (mtDNA) most frequently affecting the tRNALys gene at position m.8344A > G. Defective tRNALys severely impairs mitochondrial protein synthesis and respiratory chain when a high percentage of mutant heteroplasmy crosses the threshold for full-blown clinical phenotype. Therapy is currently limited to symptomatic management of myoclonic epilepsy, and supportive measures to counteract muscle weakness with co-factors/supplements.
Methods: We tested two therapeutic strategies to rescue mitochondrial function in cybrids and fibroblasts carrying different loads of the m.8344A > G mutation. The first strategy was aimed at inducing mitochondrial biogenesis directly, over-expressing the master regulator PGC-1α, or indirectly, through the treatment with nicotinic acid, a NAD+ precursor. The second was aimed at stimulating the removal of damaged mitochondria through prolonged rapamycin treatment.
Results: The first approach slightly increased mitochondrial protein expression and respiration in the wild type and intermediate-mutation load cells, but was ineffective in high-mutation load cell lines. This suggests that induction of mitochondrial biogenesis may not be sufficient to rescue mitochondrial dysfunction in MERRF cells with high-mutation load. The second approach, when administered chronically (4 weeks), induced a slight increase of mitochondrial respiration in fibroblasts with high-mutation load, and a significant improvement in fibroblasts with intermediate-mutation load, rescuing completely the bioenergetics defect. This effect was mediated by increased mitochondrial biogenesis, possibly related to the rapamycin-induced inhibition of the Mechanistic Target of Rapamycin Complex 1 (mTORC1) and the consequent activation of the Transcription Factor EB (TFEB).
Conclusions: Overall, our results point to rapamycin-based therapy as a promising therapeutic option for MERRF.
Keywords: MERRF; Mitochondrial DNA; Mitochondrial biogenesis; Mitochondrial dysfunction; Niacin; PGC-1α; Rapamycin; mTORC1.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no conflict of interest related to this study.
Figures
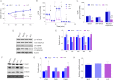
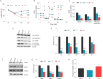


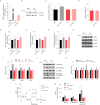
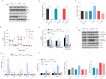
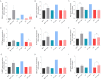
Similar articles
-
Mitochondrial impairment and synaptic dysfunction are associated with neurological defects in iPSCs-derived cortical neurons of MERRF patients.J Biomed Sci. 2023 Aug 21;30(1):70. doi: 10.1186/s12929-023-00966-8. J Biomed Sci. 2023. PMID: 37605213 Free PMC article.
-
Nuclear DNA-encoded tRNAs targeted into mitochondria can rescue a mitochondrial DNA mutation associated with the MERRF syndrome in cultured human cells.Hum Mol Genet. 2004 Oct 15;13(20):2519-34. doi: 10.1093/hmg/ddh267. Epub 2004 Aug 18. Hum Mol Genet. 2004. PMID: 15317755
-
Pathogenetic aspects of the A8344G mutation of mitochondrial DNA associated with MERRF syndrome and multiple symmetric lipomas.Muscle Nerve Suppl. 1995;3:S102-6. doi: 10.1002/mus.880181421. Muscle Nerve Suppl. 1995. PMID: 7603509 Review.
-
Distal weakness with respiratory insufficiency caused by the m.8344A > G "MERRF" mutation.Neuromuscul Disord. 2014 Jun;24(6):533-6. doi: 10.1016/j.nmd.2014.03.011. Epub 2014 Apr 1. Neuromuscul Disord. 2014. PMID: 24792523 Free PMC article.
-
[Molecular genetic analysis for myoclonus epilepsy associated with ragged-red fibers (MERRF)].Nihon Rinsho. 1993 Sep;51(9):2379-85. Nihon Rinsho. 1993. PMID: 8411716 Review. Japanese.
Cited by
-
Mitochondrial Mutations Can Alter Neuromuscular Transmission in Congenital Myasthenic Syndrome and Mitochondrial Disease.Int J Mol Sci. 2023 May 9;24(10):8505. doi: 10.3390/ijms24108505. Int J Mol Sci. 2023. PMID: 37239850 Free PMC article. Review.
-
Glutamine metabolism in diseases associated with mitochondrial dysfunction.Mol Cell Neurosci. 2023 Sep;126:103887. doi: 10.1016/j.mcn.2023.103887. Epub 2023 Aug 15. Mol Cell Neurosci. 2023. PMID: 37586651 Free PMC article. Review.
-
Progressive Myoclonus Epilepsy: A Scoping Review of Diagnostic, Phenotypic and Therapeutic Advances.Genes (Basel). 2024 Jan 27;15(2):171. doi: 10.3390/genes15020171. Genes (Basel). 2024. PMID: 38397161 Free PMC article.
References
-
- Altmann J, Büchner B, Nadaj-Pakleza A, Schäfer J, Jackson S, Lehmann D, et al. Expanded phenotypic spectrum of the m.8344A>G “MERRF” mutation: data from the German mitoNET registry. J Neurol. 2016;263(5):961–972. - PubMed
-
- Antonická H, Floryk D, Klement P, Stratilová L, Hermanská J, Houstková H, et al. Defective kinetics of cytochrome c oxidase and alteration of mitochondrial membrane potential in fibroblasts and cytoplasmic hybrid cells with the mutation for myoclonus epilepsy with ragged-red fibres ('MERRF’) at position 8344 nt. Biochem J. 1999;342(Pt 3):537–544. - PMC - PubMed
-
- Attardi G, Yoneda M, Chomyn A. Complementation and segregation behavior of disease-causing mitochondrial DNA mutations in cellular model systems. Biochim Biophys Acta Mol Basis Dis. 1995;1271(1):241–248. - PubMed
-
- Bentlage HACM, Attardi G. Relationship of genotype to phenotype in fibroblast-derived transmitochondrial cell lines carrying the 3243 mutation associated with the melas encephalomyopathy: shift towards mutant genotype and role of mtDNA copy number. Hum Mol Genet. 1996;5(2):197–205. - PubMed
-
- Berkovic SF, Carpenter S, Evans A, Karpati G, Shoubridge EA, Andermann F, et al. Myoclonus epilepsy and ragged-red fibres (MERRF). 1. A clinical, pathological, biochemical, magnetic resonance spectrographic and positron emission tomographic study. Brain. 1989;112(Pt 5):1231–1260. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

