Pseudomonas spp. Enriched in Endophytic Community of Healthy Cotton Plants Inhibit Cotton Verticillium Wilt
- PMID: 35923406
- PMCID: PMC9339998
- DOI: 10.3389/fmicb.2022.906732
Pseudomonas spp. Enriched in Endophytic Community of Healthy Cotton Plants Inhibit Cotton Verticillium Wilt
Abstract
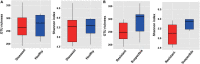
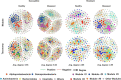
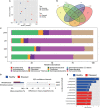
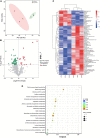
The plant microbiome plays a fundamental role in plant growth and health. However, detailed information regarding the plant endophytic microbiome during the infection period of a pathogen is largely unknown. Here, we investigated the microbial community of healthy and diseased cotton plants and the root exudate profiles of susceptible and resistant cultivars utilizing high-throughput sequencing and metabolomics. The results showed that the pathogen infection reduced bacterial diversity and significantly affected the bacterial community composition. The microbiome assembly is shaped predominantly by cultivars. The endophytic microbiome of the infected plants showed greater complexity than the healthy plants in network analysis. The results displayed that a total of 76 compounds were significantly different in the two groups, with 18 compounds showing a higher relative abundance in the resistant cultivars and 58 compounds in the susceptible cultivars. Pathway enrichment analysis showed that pathways related to plant hormone signal transduction, biosynthesis of various secondary metabolites, and biosynthesis and metabolism of amino acids were prominently altered. We also demonstrate that plants inoculated with Pseudomonas sp. strains showed increased resistance to the cotton Verticillium wilt compared with the control plants in pot experiments. Overall, it showed that the pathogen infection affected the community composition, and healthy plants displayed an enriched beneficial microbiome to combat the plant disease. These findings significantly advance our understanding of the endophytic microbiome assembly under the pathogen infection and develop microbiome-based solutions for sustainable crop production systems.
Keywords: bacterial community; beneficial microbe; cotton Verticillium wilt; endophytic; microbiome assembly.
Copyright © 2022 Zeng, Man, Dai and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Analysis of endophytic bacterial diversity in seeds of different genotypes of cotton and the suppression of Verticillium wilt pathogen infection by a synthetic microbial community.BMC Plant Biol. 2024 Apr 10;24(1):263. doi: 10.1186/s12870-024-04910-2. BMC Plant Biol. 2024. PMID: 38594616 Free PMC article.
-
Different responses of the rhizosphere microbiome to Verticillium dahliae infection in two cotton cultivars.Front Microbiol. 2023 Aug 11;14:1229454. doi: 10.3389/fmicb.2023.1229454. eCollection 2023. Front Microbiol. 2023. PMID: 37637103 Free PMC article.
-
The Endophytic Root Microbiome Is Different in Healthy and Ralstonia solanacearum-Infected Plants and Is Regulated by a Consortium Containing Beneficial Endophytic Bacteria.Microbiol Spectr. 2023 Feb 14;11(1):e0203122. doi: 10.1128/spectrum.02031-22. Epub 2022 Dec 14. Microbiol Spectr. 2023. PMID: 36515552 Free PMC article.
-
Regulatory Network of Cotton Genes in Response to Salt, Drought and Wilt Diseases (Verticillium and Fusarium): Progress and Perspective.Front Plant Sci. 2021 Nov 29;12:759245. doi: 10.3389/fpls.2021.759245. eCollection 2021. Front Plant Sci. 2021. PMID: 34912357 Free PMC article. Review.
-
Insights to Gossypium defense response against Verticillium dahliae: the Cotton Cancer.Funct Integr Genomics. 2023 May 1;23(2):142. doi: 10.1007/s10142-023-01065-5. Funct Integr Genomics. 2023. PMID: 37121989 Review.
Cited by
-
Grapevines escaping trunk diseases in New Zealand vineyards have a distinct microbiome structure.Front Microbiol. 2023 Aug 23;14:1231832. doi: 10.3389/fmicb.2023.1231832. eCollection 2023. Front Microbiol. 2023. PMID: 37680529 Free PMC article.
-
Next-generation sequencing-based comparative mapping and culture-based screening of bacterial rhizobiome in Phytophthora capsici-resistant and susceptible Piper species.Front Microbiol. 2024 Sep 25;15:1458454. doi: 10.3389/fmicb.2024.1458454. eCollection 2024. Front Microbiol. 2024. PMID: 39403087 Free PMC article.
-
Analysis of endophytic bacterial diversity in seeds of different genotypes of cotton and the suppression of Verticillium wilt pathogen infection by a synthetic microbial community.BMC Plant Biol. 2024 Apr 10;24(1):263. doi: 10.1186/s12870-024-04910-2. BMC Plant Biol. 2024. PMID: 38594616 Free PMC article.
-
Complete genome sequence data of Pseudomonas nitroreducens L4, an endophyte isolated from cotton plants.Data Brief. 2024 Jun 13;55:110639. doi: 10.1016/j.dib.2024.110639. eCollection 2024 Aug. Data Brief. 2024. PMID: 39022698 Free PMC article.
-
The impact of filamentous plant pathogens on the host microbiota.BMC Biol. 2024 Aug 15;22(1):175. doi: 10.1186/s12915-024-01965-3. BMC Biol. 2024. PMID: 39148076 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

