Crystal structures of glutamyl-tRNA synthetase from Elizabethkingia anopheles and E. meningosepticum
- PMID: 35924598
- PMCID: PMC9350836
- DOI: 10.1107/S2053230X22007555
Crystal structures of glutamyl-tRNA synthetase from Elizabethkingia anopheles and E. meningosepticum
Abstract
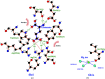
Elizabethkingia bacteria are globally emerging pathogens that cause opportunistic and nosocomial infections, with up to 40% mortality among the immunocompromised. Elizabethkingia species are in the pipeline of organisms for high-throughput structural analysis at the Seattle Structural Genomics Center for Infectious Disease (SSGCID). These efforts include the structure-function analysis of potential therapeutic targets. Glutamyl-tRNA synthetase (GluRS) is essential for tRNA aminoacylation and is under investigation as a bacterial drug target. The SSGCID produced, crystallized and determined high-resolution structures of GluRS from E. meningosepticum (EmGluRS) and E. anopheles (EaGluRS). EmGluRS was co-crystallized with glutamate, while EaGluRS is an apo structure. EmGluRS shares ∼97% sequence identity with EaGluRS but less than 39% sequence identity with any other structure in the Protein Data Bank. EmGluRS and EaGluRS have the prototypical bacterial GluRS topology. EmGluRS and EaGluRS have similar binding sites and tertiary structures to other bacterial GluRSs that are promising drug targets. These structural similarities can be exploited for drug discovery.
Keywords: Elizabethkingia anopheles; Elizabethkingia meningosepticum; Seattle Structural Genomics Center for Infectious Disease; emerging infectious diseases; glutamyl-tRNA synthetases; infectious diseases; undergraduate education and training.
open access.
Figures


Similar articles
-
Crystal structure of glutamyl-tRNA synthetase from Helicobacter pylori.Acta Crystallogr F Struct Biol Commun. 2024 Dec 1;80(Pt 12):335-40. doi: 10.1107/S2053230X24011099. Online ahead of print. Acta Crystallogr F Struct Biol Commun. 2024. PMID: 39601417 Free PMC article.
-
Signatures of tRNAGlx-specificity in proteobacterial glutamyl-tRNA synthetases.Proteins. 2025 Jan;93(1):241-254. doi: 10.1002/prot.26634. Epub 2023 Nov 12. Proteins. 2025. PMID: 37953434
-
A C-truncated glutamyl-tRNA synthetase specific for tRNA(Glu) is stimulated by its free complementary distal domain: mechanistic and evolutionary implications.Biochemistry. 2009 Jun 30;48(25):6012-21. doi: 10.1021/bi801690f. Biochemistry. 2009. PMID: 19496540
-
Glutamyl-tRNA sythetase.Biol Chem. 1997 Nov;378(11):1313-29. Biol Chem. 1997. PMID: 9426192 Review.
-
Substrate selection by aminoacyl-tRNA synthetases.Nucleic Acids Symp Ser. 1995;(33):40-2. Nucleic Acids Symp Ser. 1995. PMID: 8643392 Review.
Cited by
-
Using resources generated by the Seattle Structural Genomics Center for Infectious Disease (SSGCID) for training early career researchers.Acta Crystallogr F Struct Biol Commun. 2025 Jun 1;81(Pt 6):222-225. doi: 10.1107/S2053230X25003930. Epub 2025 May 6. Acta Crystallogr F Struct Biol Commun. 2025. PMID: 40439320 Free PMC article.
-
Crystal structure of glutamyl-tRNA synthetase from Helicobacter pylori.Acta Crystallogr F Struct Biol Commun. 2024 Dec 1;80(Pt 12):335-40. doi: 10.1107/S2053230X24011099. Online ahead of print. Acta Crystallogr F Struct Biol Commun. 2024. PMID: 39601417 Free PMC article.
-
Architecture of Pseudomonas aeruginosa glutamyl-tRNA synthetase defines a subfamily of dimeric class Ib aminoacyl-tRNA synthetases.Proc Natl Acad Sci U S A. 2025 May 13;122(19):e2504757122. doi: 10.1073/pnas.2504757122. Epub 2025 May 9. Proc Natl Acad Sci U S A. 2025. PMID: 40343997
-
Structures of Brucella ovis leucine-, isoleucine-, valine-, threonine- and alanine-binding protein reveal a conformationally flexible peptide-binding cavity.Acta Crystallogr F Struct Biol Commun. 2024 Sep 1;80(Pt 9):193-199. doi: 10.1107/S2053230X24007027. Epub 2024 Aug 23. Acta Crystallogr F Struct Biol Commun. 2024. PMID: 39177244 Free PMC article.
References
-
- Adams, P. D., Afonine, P. V., Bunkóczi, G., Chen, V. B., Echols, N., Headd, J. J., Hung, L.-W., Jain, S., Kapral, G. J., Grosse Kunstleve, R. W., McCoy, A. J., Moriarty, N. W., Oeffner, R. D., Read, R. J., Richardson, D. C., Richardson, J. S., Terwilliger, T. C. & Zwart, P. H. (2011). Methods, 55, 94–106. - PMC - PubMed
-
- Aravind, L., Anantharaman, V. & Koonin, E. V. (2002). Proteins, 48, 1–14. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

