Characterization of a Novel RNA Virus Causing Massive Mortality in Yellow Catfish, Pelteobagrus fulvidraco, as an Emerging Genus in Caliciviridae (Picornavirales)
- PMID: 35924844
- PMCID: PMC9431444
- DOI: 10.1128/spectrum.00624-22
Characterization of a Novel RNA Virus Causing Massive Mortality in Yellow Catfish, Pelteobagrus fulvidraco, as an Emerging Genus in Caliciviridae (Picornavirales)
Abstract
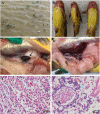
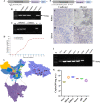
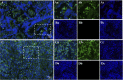
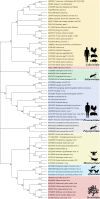
An emerging disease in farmed yellow catfish (Pelteobagrus fulvidraco) causing massive mortality broke out in 2020 in Hubei, China. Histopathological examination indicated significant changes in kidneys and spleens of diseased fish. Electron microscopy revealed large numbers of viral particles in the kidneys and spleens. These particles were spherical with a diameter of approximately 35 nm. By using RNA sequencing and rapid identification of cDNA ends, the full nucleotide sequence of the virus was identified. The viral genome comprises 7,432 bp and contains three open reading frames sharing no nucleotide sequence similarity with other viruses; however, the amino acid sequence partially matched that of the nonstructural (NS) proteins from viruses in the order Picornavirales. Combined with the phylogenetic analysis, the conserved amino acid motifs and the domains of the viral genome predict a genome order typical of a calicivirus. Therefore, this virus was tentatively named yellow catfish calicivirus (YcCV). Cell culture showed that YcCV could cause a cytopathic effect in the channel catfish kidney cell line (CCK) at early passages. In artificial infection, this virus could infect healthy yellow catfish and led to clinical symptoms similar to those that occurred naturally. In situ hybridization analysis detected positive signals of the virus in kidney, spleen, liver, heart, and gill tissues of diseased fish. This study represents the first report of calicivirus infection in yellow catfish and provides a solid basis for future studies on the control of this viral disease. IMPORTANCE Caliciviruses are rapidly evolving viruses that cause pandemic outbreaks associated with significant morbidity and mortality globally. A novel calicivirus identified from yellow catfish also causes substantial mortality. Using an RNA sequencing (RNA-seq) and rapid amplification of cDNA ends (RACE) method, the full nucleotide sequence was identified and characterized, and this virus was tentatively named yellow catfish calicivirus (YcCV). A nucleotide sequence similarity search found no match with other viruses, and an amino acid sequence comparison indicated approximately 23.3% amino acid homology with the viruses in the order Picornavirales. These findings may represent a new avenue to explain virus evolution and suggest a need to further study the pathogenesis of calicivirus and characterize possible interactions among interspecific viruses in the aquaculture environment.
Keywords: Calicivirus; Pelteobagrus fulvidraco; Picornavirus; characterization; phylogenetic analysis; yellow catfish.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
RNA-Sequencing Analysis of the Viral Community in Yellow Catfish (Pelteobagrus fulvidraco) in the Upper Reaches of the Yangtze River.Animals (Basel). 2024 Nov 25;14(23):3386. doi: 10.3390/ani14233386. Animals (Basel). 2024. PMID: 39682352 Free PMC article.
-
Natural Co-Infections of Aeromonas veronii and Yellow Catfish Calicivirus (YcCV) in Ascites Disease Outbreaks in Cultured Yellow Catfish: An Emerging Fish Disease in China.Animals (Basel). 2024 Nov 15;14(22):3289. doi: 10.3390/ani14223289. Animals (Basel). 2024. PMID: 39595340 Free PMC article.
-
Molecular identification and expression analysis of a goose-type lysozyme (LysG) gene in yellow catfish Pelteobagrus fulvidraco.Fish Shellfish Immunol. 2016 Nov;58:423-428. doi: 10.1016/j.fsi.2016.09.034. Epub 2016 Sep 16. Fish Shellfish Immunol. 2016. PMID: 27645907
-
Functions of the 5' and 3' ends of calicivirus genomes.Virus Res. 2015 Aug 3;206:134-43. doi: 10.1016/j.virusres.2015.02.002. Epub 2015 Feb 9. Virus Res. 2015. PMID: 25678268 Free PMC article. Review.
-
Calicivirus RNA-Dependent RNA Polymerases: Evolution, Structure, Protein Dynamics, and Function.Front Microbiol. 2019 Jun 6;10:1280. doi: 10.3389/fmicb.2019.01280. eCollection 2019. Front Microbiol. 2019. PMID: 31244803 Free PMC article. Review.
Cited by
-
Diversity, evolution, and emergence of fish viruses.J Virol. 2024 Jun 13;98(6):e0011824. doi: 10.1128/jvi.00118-24. Epub 2024 May 24. J Virol. 2024. PMID: 38785422 Free PMC article. Review.
-
Viromics Reveals the High Diversity of Viruses from Fishes of the Tibet Highland.Microbiol Spectr. 2023 Jun 15;11(3):e0094623. doi: 10.1128/spectrum.00946-23. Epub 2023 May 23. Microbiol Spectr. 2023. PMID: 37219423 Free PMC article.
-
Spatiotemporal Dynamics of Pro-Inflammatory Mediator Expression in Pelteobagrus vachelli During Ichthyophthiriasis: A 40-Day Longitudinal Study of IL-1β, IL-6, and SAA.Animals (Basel). 2025 May 28;15(11):1577. doi: 10.3390/ani15111577. Animals (Basel). 2025. PMID: 40509043 Free PMC article.
-
RNA-Sequencing Analysis of the Viral Community in Yellow Catfish (Pelteobagrus fulvidraco) in the Upper Reaches of the Yangtze River.Animals (Basel). 2024 Nov 25;14(23):3386. doi: 10.3390/ani14233386. Animals (Basel). 2024. PMID: 39682352 Free PMC article.
References
-
- Ministry of Agriculture, China. 2021. China fisheries statistical yearbook. China Agriculture Publisher, Beijing, China.
-
- Ye SG, Li H, Qiao G, Li ZS. 2009. First case of Edwardsiella ictaluri infection in China farmed yellow catfish Pelteobagrus fulvidraco. Aquaculture 292:6–10. doi:10.1016/j.aquaculture.2009.03.036. - DOI
-
- Kim JD, Park SW. 2015. Edwardsiella ictaluri infection in cultured yellow catfish Pelteobagrus fulvidraco fingerlings in Korea. Korean J Fish Aquat Sci 48:725–730. doi:10.5657/KFAS.2015.0725. - DOI
-
- Zhou Y, Jiang N, Zeng J, Fan YD, Liu WZ, Si KG, Zeng LB. 2019. Isolation and identification of pathogenic bacterium from ascites disease of yellow catfish, Pelteobagrus fulvidraco. Chin Fish Qual Stand 9:2095.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

