Genetic Differentiation and Migration Fluxes of Viruses from Melon Crops and Crop Edge Weeds
- PMID: 35924924
- PMCID: PMC9400485
- DOI: 10.1128/jvi.00421-22
Genetic Differentiation and Migration Fluxes of Viruses from Melon Crops and Crop Edge Weeds
Abstract
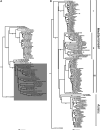
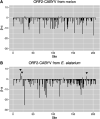
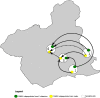
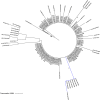
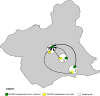
Weeds surrounding crops may act as alternative hosts, playing important epidemiological roles as virus reservoirs and impacting virus evolution. We used high-throughput sequencing to identify viruses in Spanish melon crops and plants belonging to three pluriannual weed species, Ecballium elaterium, Malva sylvestris, and Solanum nigrum, sampled at the edges of the crops. Melon and E. elaterium, both belonging to the family Cucurbitaceae, shared three virus species, whereas there was no virus species overlap between melon and the other two weeds. The diversity of cucurbit aphid-borne yellows virus (CABYV) and tomato leaf curl New Delhi virus (ToLCNDV), both in melon and E. elaterium, was further studied by amplicon sequencing. Phylogenetic and population genetics analyses showed that the CABYV population was structured by the host, identifying three sites in the CABYV RNA-dependent RNA polymerase under positive selection, perhaps reflecting host adaptation. The ToLCNDV population was much less diverse than the CABYV one, likely as a consequence of the relatively recent introduction of ToLCNDV in Spain. In spite of its low diversity, we identified geographical but no host differentiation for ToLCNDV. Potential virus migration fluxes between E. elaterium and melon plants were also analyzed. For CABYV, no evidence of migration between the populations of the two hosts was found, whereas important fluxes were identified between geographically distant subpopulations for each host. For ToLCNDV, in contrast, evidence of migration from melon to E. elaterium was found, but not the other way around. IMPORTANCE It has been reported that about half of the emerging diseases affecting plants are caused by viruses. Alternative hosts often play critical roles in virus emergence as virus reservoirs, bridging host species that are otherwise unconnected and/or favoring virus diversification. In spite of this, the viromes of potential alternative hosts remain largely unexplored. In the case of crops, pluriannual weeds at the crop edges may play these roles. Here, we took advantage of the power of high-throughput sequencing to characterize the viromes of three weed species frequently found at the edges of melon crops. We identified three viruses shared by melon and the cucurbit weed, with two of them being epidemiologically relevant for melon crops. Further genetic analyses showed that these two viruses had contrasting patterns of diversification and migration, providing an interesting example on the role that weeds may play in the ecology and evolution of viruses affecting crops.
Keywords: CABYV; CCYV; CmEV; ToLCNDV; genetic differentiation; high-throughput sequencing; migration fluxes; virus reservoirs.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
In-depth study of tomato and weed viromes reveals undiscovered plant virus diversity in an agroecosystem.Microbiome. 2023 Mar 28;11(1):60. doi: 10.1186/s40168-023-01500-6. Microbiome. 2023. PMID: 36973750 Free PMC article.
-
Home treatment for mental health problems: a systematic review.Health Technol Assess. 2001;5(15):1-139. doi: 10.3310/hta5150. Health Technol Assess. 2001. PMID: 11532236
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Intravenous magnesium sulphate and sotalol for prevention of atrial fibrillation after coronary artery bypass surgery: a systematic review and economic evaluation.Health Technol Assess. 2008 Jun;12(28):iii-iv, ix-95. doi: 10.3310/hta12280. Health Technol Assess. 2008. PMID: 18547499
-
Drugs for preventing postoperative nausea and vomiting in adults after general anaesthesia: a network meta-analysis.Cochrane Database Syst Rev. 2020 Oct 19;10(10):CD012859. doi: 10.1002/14651858.CD012859.pub2. Cochrane Database Syst Rev. 2020. PMID: 33075160 Free PMC article.
Cited by
-
Occurrence, Distribution, and Management of Aphid-Transmitted Viruses in Cucurbits in Spain.Pathogens. 2023 Mar 7;12(3):422. doi: 10.3390/pathogens12030422. Pathogens. 2023. PMID: 36986344 Free PMC article. Review.
-
Overview of research on virus-resistant breeding of melon.Front Plant Sci. 2024 Dec 12;15:1500246. doi: 10.3389/fpls.2024.1500246. eCollection 2024. Front Plant Sci. 2024. PMID: 39726431 Free PMC article. Review.
-
Benchmarking of virome metagenomic analysis approaches using a large, 60+ members, viral synthetic community.J Virol. 2023 Nov 30;97(11):e0130023. doi: 10.1128/jvi.01300-23. Epub 2023 Oct 27. J Virol. 2023. PMID: 37888981 Free PMC article.
-
Carrot populations in France and Spain host a complex virome rich in previously uncharacterized viruses.PLoS One. 2023 Aug 16;18(8):e0290108. doi: 10.1371/journal.pone.0290108. eCollection 2023. PLoS One. 2023. PMID: 37585477 Free PMC article.
-
New Weed Hosts for Tomato Brown Rugose Fruit Virus in Wild Mediterranean Vegetation.Plants (Basel). 2022 Sep 1;11(17):2287. doi: 10.3390/plants11172287. Plants (Basel). 2022. PMID: 36079668 Free PMC article.
References
-
- Mutuku JM, Wamonje FO, Mukeshimana G, Njuguna J, Wamalwa M, Choi S-K, Tungadi T, Djikeng A, Kelly K, Domelevo Entfellner J-B, Ghimire SR, Mignouna HD, Carr JP, Harvey JJW. 2018. Metagenomic analysis of plant virus occurrence in common bean (Phaseolus vulgaris) in Central Kenya. Front Microbiol 9:2939. 10.3389/fmicb.2018.02939. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources

