Towards systematic exploration of chemical space: building the fragment library module in molecular property diagnostic suite
- PMID: 35925528
- PMCID: PMC9362107
- DOI: 10.1007/s11030-022-10506-5
Towards systematic exploration of chemical space: building the fragment library module in molecular property diagnostic suite
Abstract
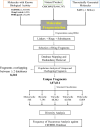
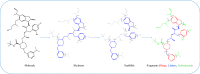
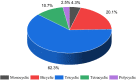
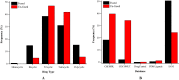
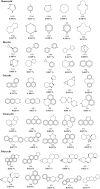
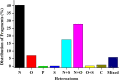
A fragment-based drug discovery (FBDD) approach has traditionally been of utmost significance in drug design studies. It allows the exploration of large chemical space to find novel scaffolds and chemotypes which can be improved into selective inhibitors with good affinity. In the current work, several public domain chemical libraries (ChEMBL, DrugCentral, PDB ligands, COCONUT, and SAVI) comprising bioactive and virtual molecules were retrieved to develop a fragment library. A systematic fragmentation method that breaks a given molecule into rings, linkers, and substituents was used to cleave the molecules and the fragments were analyzed. Further, only the ring framework was taken into the consideration to develop a fragment library that consists of a total number of 107,614 unique fragments. This set represents a rich diverse structure framework that covers a wide variety of yet-to-be-explored fragments for a wide range of small molecule-based applications. This fragment library is an integral part of the molecular property diagnostic suite (MPDS) suite that can be used with other modeling and informatics methods for FBDD approaches. The fragment library module of MPDS can be accessed at http://mpds.neist.res.in:8085 .
Keywords: Drug discovery; Fragment library; Fragment space; MPDS.
© 2022. The Author(s), under exclusive licence to Springer Nature Switzerland AG.
Conflict of interest statement
The authors have no conflict of interest.
Figures
References
-
- Gaur AS, Bhardwaj A, Sharma A, John L, Vivek MR, Tripathi N, Bharatam PV, Kumar R, Janardhan S, Mori A, Banerji A, Lynn AM, Hemrom AJ, Passi A, Singh A, Kumar A, Muvva C, Madhuri C, Choudhury C, Kumar AD, Pandit D, Bharti DR, Kumar D, Singam AE, Raghava GPS, Sailaja H, Jangra H, Raithatha K, Tanneeru K, Chaudhary K, Karthikeyan M, Prasanthi M, Kumar N, Yedukondalu N, Rajput NK, Saranya PS, Narang P, Dutta P, Krishnan RV, Sharma R, Srinithi R, Mishra R, Hemasri S, Singh S, Venkatesan S, Kumar S, Jaleel UCA, Khedkar V, Joshi Y, Sastry GN. Assessing therapeutic potential of molecules: molecular property diagnostic suite for tuberculosis (MPDSTB) J Chem Sci. 2017;129:515. doi: 10.1007/s12039-017-1268-4. - DOI
-
- Nagamani S, Gaur AS, Tanneeru K, Muneeswaran G, Madugula SS, MPDS Consortium, Druzhilovskiy D, Poroikov VV, Sastry GN Molecular property diagnostic suite (MPDS): development of disease-specific open-source web portals for drug discovery. SAR QSAR Environ Res. 2017;11:913–926. doi: 10.1080/1062936X.2017.1402819. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

