Identifying signatures of natural selection in Indian populations
- PMID: 35925921
- PMCID: PMC9352006
- DOI: 10.1371/journal.pone.0271767
Identifying signatures of natural selection in Indian populations
Abstract
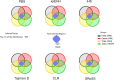
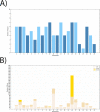
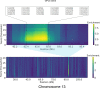
In this study, we present the results of a genome-wide scan for signatures of positive selection using data from four tribal groups (Kokana, Warli, Bhil, and Pawara) and two caste groups (Deshastha Brahmin and Kunbi Maratha) from West of the Maharashtra State In India, as well as two samples of South Asian ancestry from the 1KG project (Gujarati Indian from Houston, Texas and Indian Telugu from UK). We used an outlier approach based on different statistics, including PBS, xpEHH, iHS, CLR, Tajima's D, as well as two recently developed methods: Graph-aware Retrieval of Selective Sweeps (GRoSS) and Ascertained Sequentially Markovian Coalescent (ASMC). In order to minimize the risk of false positives, we selected regions that are outliers in all the samples included in the study using more than one method. We identified putative selection signals in 107 regions encompassing 434 genes. Many of the regions overlap with only one gene. The signals observed using microarray-based data are very consistent with our analyses using high-coverage sequencing data, as well as those identified with a novel coalescence-based method (ASMC). Importantly, at least 24 of these genomic regions have been identified in previous selection scans in South Asian populations or in other population groups. Our study highlights genomic regions that may have played a role in the adaptation of anatomically modern humans to novel environmental conditions after the out of Africa migration.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

