Sib1, Sib2, and Sib3 proteins are required for ferrichrome-mediated cross-feeding interaction between Schizosaccharomyces pombe and Saccharomyces cerevisiae
- PMID: 35928155
- PMCID: PMC9344042
- DOI: 10.3389/fmicb.2022.962853
Sib1, Sib2, and Sib3 proteins are required for ferrichrome-mediated cross-feeding interaction between Schizosaccharomyces pombe and Saccharomyces cerevisiae
Abstract
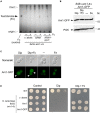
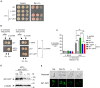
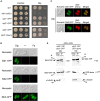
Although Saccharomyces cerevisiae is unable to produce siderophores, this fungal organism can assimilate iron bound to the hydroxamate-type siderophore ferrichrome (Fc) produced and secreted by other microbes. Fc can enter S. cerevisiae cells via Arn1. Unlike S. cerevisiae, Schizosaccharomyces pombe synthesizes and secretes Fc. The sib1 + and sib2 + genes encode, respectively, a Fc synthetase and an ornithine-N5-oxygenase, which are required for Fc production. When both genes were expressed in S. pombe, cross-feeding experiments revealed that S. cerevisiae fet3Δ arn1-4Δ cells expressing Arn1 could grow in the vicinity of S. pombe under low-iron conditions. In contrast, deletion of sib1 + and sib2 + produced a defect in the ability of S. pombe to keep S. cerevisiae cells alive when Fc is used as the sole source of iron. Further analysis identified a gene designated sib3 + that encodes an N5-transacetylase required for Fc production in S. pombe. The sib3Δ mutant strain exhibited a severe growth defect in iron-poor media, and it was unable to promote Fc-dependent growth of S. cerevisiae cells. Microscopic analyses of S. pombe cells expressing a functional Sib3-GFP protein revealed that Sib3 was localized throughout the cells, with a proportion of Sib3 being colocalized with Sib1 and Sib2 within the cytosol. Collectively, these results describe the first example of a one-way cross-feeding interaction, with S. pombe providing Fc that enables S. cerevisiae to grow when Fc is used as the sole source of iron.
Keywords: budding yeast; cross-feeding; ferrichrome; fission yeast; siderophore.
Copyright © 2022 Brault, Mbuya and Labbé.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

