Identification of hub genes and their correlation with immune infiltration in coronary artery disease through bioinformatics and machine learning methods
- PMID: 35928610
- PMCID: PMC9344404
- DOI: 10.21037/jtd-22-632
Identification of hub genes and their correlation with immune infiltration in coronary artery disease through bioinformatics and machine learning methods
Abstract
Background: Coronary artery disease (CAD) is a multifactorial disease and its pathogenesis remains unclear. We aimed to explore the optimal feature genes (OFGs) for CAD and to investigate the function of immune cell infiltration of CAD. It will be helpful for better understanding of the pathogenesis and the development of genetic prediction of CAD.
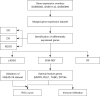
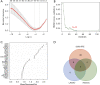
Methods: Datasets related to CAD were obtained from the Gene Expression Omnibus (GEO) database. Cases from the datasets met diagnostic criteria including clinical symptoms, electrocardiographic (ECG) and angiographic evidence. We identified differentially expressed genes (DEGs) and conducted functional enrichment analysis. OFGs were obtained from the least absolute shrinkage and selection operator (LASSO) algorithm, support vector machine recursive feature elimination (SVM-RFE) algorithm, and random forest (RF) algorithm. CIBERSORT was used to compare immune infiltration between CAD patients and normal controls, and the correlation between OFGs and immune cells was analyzed.
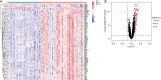
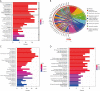
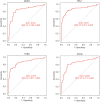
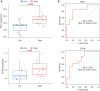
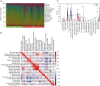
Results: DEGs were involved in the interleukin (IL)-17 signaling pathway, nuclear factor (NF)-kappa B signaling pathway, and tumor necrosis factor (TNF) signaling pathway. Gene Ontology (GO) analysis revealed DEGs were enriched in lipopolysaccharide (LPS), tertiary granule, and pattern recognition receptor activity. Disease Ontology (DO) analysis suggested DEGs were enriched in lung disease, arteriosclerotic cardiovascular disease (CVD). Matrix metalloproteinase 9 (MMP9), Pellino E3 ubiquitin protein ligase 1 (PELI1), thrombomodulin (THBD), and zinc finger protein 36 (ZFP36) were screened by the intersection of OFGs obtained from LASSO, SVM-REF, and RF algorithms. CAD patients had a lower proportion of memory B cells (P=0.019), CD8 T cells (P<0.001), resting memory CD4 T cells (P<0.001), regulatory T cells (P=0.028), and gamma delta T cells (P<0.001) than normal controls, while the proportion of activated memory CD4 T cells (P=0.014), resting natural killer (NK) cells (P<0.001), monocytes (P<0.001), M0 macrophages (P=0.023), activated mast cells (P<0.001), and neutrophils (P<0.001) in CAD patients were higher than normal controls. MMP9, PELI1, THBD, and ZFP36 were correlated with immune cells.
Conclusions: MMP9, PELI1, THBD, and ZFP36 may be predicted biomarkers for CAD. The OFGs and association between OFGs and immune infiltration may provide potential biomarkers for CAD prediction along with the better assessment of the disease.
Keywords: Coronary artery disease (CAD); bioinformatics analysis; immune infiltration; machine learning (ML); optimal feature genes (OFGs).
2022 Journal of Thoracic Disease. All rights reserved.
Conflict of interest statement
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jtd.amegroups.com/article/view/10.21037/jtd-22-632/coif). The authors have no conflicts of interest to declare.
Figures

Similar articles
-
Immune-related potential biomarkers and therapeutic targets in coronary artery disease.Front Cardiovasc Med. 2023 Jan 6;9:1055422. doi: 10.3389/fcvm.2022.1055422. eCollection 2022. Front Cardiovasc Med. 2023. PMID: 36684561 Free PMC article.
-
Bioinformatic Analysis and Machine Learning Methods in Neonatal Sepsis: Identification of Biomarkers and Immune Infiltration.Biomedicines. 2023 Jun 28;11(7):1853. doi: 10.3390/biomedicines11071853. Biomedicines. 2023. PMID: 37509492 Free PMC article.
-
Identification of a novel immune infiltration-related gene signature, MCEMP1, for coronary artery disease.PeerJ. 2024 Sep 25;12:e18135. doi: 10.7717/peerj.18135. eCollection 2024. PeerJ. 2024. PMID: 39346078 Free PMC article.
-
JUN and ATF3 in Gout: Ferroptosis-related potential diagnostic biomarkers.Heliyon. 2024 Oct 30;10(22):e39957. doi: 10.1016/j.heliyon.2024.e39957. eCollection 2024 Nov 30. Heliyon. 2024. PMID: 39619595 Free PMC article. Review.
-
Integrated identification of key immune related genes and patterns of immune infiltration in calcified aortic valvular disease: A network based meta-analysis.Front Genet. 2022 Sep 21;13:971808. doi: 10.3389/fgene.2022.971808. eCollection 2022. Front Genet. 2022. PMID: 36212153 Free PMC article.
Cited by
-
Exploring the association between rosacea and acne by integrated bioinformatics analysis.Sci Rep. 2024 Feb 6;14(1):3065. doi: 10.1038/s41598-024-53453-x. Sci Rep. 2024. PMID: 38321132 Free PMC article.
-
Identification of novel biomarkers and immune infiltration features of recurrent pregnancy loss by machine learning.Sci Rep. 2023 Jul 3;13(1):10751. doi: 10.1038/s41598-023-38046-4. Sci Rep. 2023. PMID: 37400532 Free PMC article.
-
Early heart disease prediction using feature engineering and machine learning algorithms.Heliyon. 2024 Oct 1;10(19):e38731. doi: 10.1016/j.heliyon.2024.e38731. eCollection 2024 Oct 15. Heliyon. 2024. PMID: 39397946 Free PMC article.
-
Biology of Pellino1: a potential therapeutic target for inflammation in diseases and cancers.Front Immunol. 2023 Dec 18;14:1292022. doi: 10.3389/fimmu.2023.1292022. eCollection 2023. Front Immunol. 2023. PMID: 38179042 Free PMC article. Review.
-
Identification of hub biomarkers in coronary artery disease patients using machine learning and bioinformatic analyses.Sci Rep. 2025 May 18;15(1):17244. doi: 10.1038/s41598-025-02123-7. Sci Rep. 2025. PMID: 40383719 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
