Putative transcription antiterminator RfaH contributes to Erwinia amylovora virulence
- PMID: 35929143
- PMCID: PMC9562583
- DOI: 10.1111/mpp.13254
Putative transcription antiterminator RfaH contributes to Erwinia amylovora virulence
Abstract
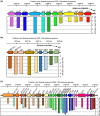
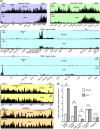
The gram-negative bacterium Erwinia amylovora causes fire blight disease of apple and pear trees. The exopolysaccharide amylovoran and lipopolysaccharides are essential E. amylovora virulence factors. Production of amylovoran and lipopolysaccharide is specified in part by genes that are members of long operons. Here, we show that full virulence of E. amylovora in apple fruitlets and tree shoots depends on the predicted transcription antiterminator RfaH. RfaH reduces pausing in the production of long transcripts having an operon polarity suppressor regulatory element within their promoter region. In E. amylovora, only the amylovoran operon and a lipopolysaccharide operon have such regulatory elements within their promoter regions and in the correct orientation. These operons showed dramatically increased polarity in the ΔrfaH mutant compared to the wild type as determined by RNA sequencing. Amylovoran and lipopolysaccharide production in vitro was reduced in rfaH mutants compared to the wild type, which probably contributes to the rfaH mutant virulence phenotype. Furthermore, type VI secretion cluster 1, which contributes to E. amylovora virulence, showed reduced expression in ΔrfaH compared to the wild type, although without an increase in polarity. The data suggest that E. amylovora RfaH directly, specifically, and exclusively suppresses operon polarity in the amylovoran operon and a lipopolysaccharide operon.
Keywords: apple; fire blight; operon; pear; type VI secretion.
© 2022 The Authors. Molecular Plant Pathology published by British Society for Plant Pathology and John Wiley & Sons Ltd.
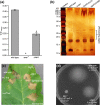
Figures

Similar articles
-
Virulence Genetics of an Erwinia amylovora Putative Polysaccharide Transporter Family Member.J Bacteriol. 2020 Oct 22;202(22):e00390-20. doi: 10.1128/JB.00390-20. Print 2020 Oct 22. J Bacteriol. 2020. PMID: 32839177 Free PMC article.
-
Genome-wide identification of genes regulated by the Rcs phosphorelay system in Erwinia amylovora.Mol Plant Microbe Interact. 2012 Jan;25(1):6-17. doi: 10.1094/MPMI-08-11-0207. Mol Plant Microbe Interact. 2012. PMID: 21936662
-
AmyR is a novel negative regulator of amylovoran production in Erwinia amylovora.PLoS One. 2012;7(9):e45038. doi: 10.1371/journal.pone.0045038. Epub 2012 Sep 18. PLoS One. 2012. PMID: 23028751 Free PMC article.
-
Virulence Factors of Erwinia amylovora: A Review.Int J Mol Sci. 2015 Jun 5;16(6):12836-54. doi: 10.3390/ijms160612836. Int J Mol Sci. 2015. PMID: 26057748 Free PMC article. Review.
-
Molecular genetics of Erwinia amylovora involved in the development of fire blight.FEMS Microbiol Lett. 2005 Dec 15;253(2):185-92. doi: 10.1016/j.femsle.2005.09.051. Epub 2005 Oct 13. FEMS Microbiol Lett. 2005. PMID: 16253442 Review.
Cited by
-
Metamorphic proteins under a computational microscope: Lessons from a fold-switching RfaH protein.Comput Struct Biotechnol J. 2022 Oct 21;20:5824-5837. doi: 10.1016/j.csbj.2022.10.024. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 36382197 Free PMC article. Review.
References
-
- Artsimovitch, I. & Landick, R. (2002) The transcriptional regulator RfaH stimulates RNA chain synthesis after recruitment to elongation complexes by the exposed nontemplate DNA strand. Cell, 109, 193–203. - PubMed
-
- Bailey, M.J.A. , Hughes, C. & Koronakis, V. (1996) Increased distal gene transcription by the elongation factor RfaH, a specialized homologue of NusG. Molecular Microbiology, 22, 729–737. - PubMed
-
- Bailey, M.J.A. , Hughes, C. & Koronakis, V. (1997) RfaH and the ops element, components of a novel system controlling bacterial transcription elongation. Molecular Microbiology, 26, 845–851. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

