Toward protein NMR at physiological concentrations by hyperpolarized water-Finding and mapping uncharted conformational spaces
- PMID: 35930648
- PMCID: PMC9355353
- DOI: 10.1126/sciadv.abq5179
Toward protein NMR at physiological concentrations by hyperpolarized water-Finding and mapping uncharted conformational spaces
Abstract
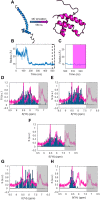
Nuclear magnetic resonance (NMR) spectroscopy is a key method for determining the structural dynamics of proteins in their native solution state. However, the low sensitivity of NMR typically necessitates nonphysiologically high sample concentrations, which often limit the relevance of the recorded data. We show how to use hyperpolarized water by dissolution dynamic nuclear polarization (DDNP) to acquire protein spectra at concentrations of 1 μM within seconds and with a high signal-to-noise ratio. The importance of approaching physiological concentrations is demonstrated for the vital MYC-associated factor X, which we show to switch conformations when diluted. While in vitro conditions lead to a population of the well-documented dimer, concentrations lowered by more than two orders of magnitude entail dimer dissociation and formation of a globularly folded monomer. We identified this structure by integrating DDNP with computational techniques to overcome the often-encountered constraint of DDNP of limited structural information provided by the typically detected one-dimensional spectra.
Figures

Similar articles
-
Nuclear Overhauser spectroscopy in hyperpolarized water - chemical vs. magnetic exchange.Chem Commun (Camb). 2022 Oct 18;58(83):11661-11664. doi: 10.1039/d2cc03735a. Chem Commun (Camb). 2022. PMID: 36169286 Free PMC article.
-
Inversion of Hyperpolarized 13C NMR Signals through Cross-Correlated Cross-Relaxation in Dissolution DNP Experiments.J Phys Chem B. 2022 Jun 23;126(24):4599-4610. doi: 10.1021/acs.jpcb.2c03375. Epub 2022 Jun 8. J Phys Chem B. 2022. PMID: 35675502 Free PMC article.
-
Micromolar Concentration Affinity Study on a Benchtop NMR Spectrometer with Secondary 13C Labeled Hyperpolarized Ligands.ACS Omega. 2025 Jan 24;10(4):3332-3337. doi: 10.1021/acsomega.4c05101. eCollection 2025 Feb 4. ACS Omega. 2025. PMID: 39926548 Free PMC article.
-
Missing Pieces in Structure Puzzles: How Hyperpolarized NMR Spectroscopy Can Complement Structural Biology and Biochemistry.Chembiochem. 2023 Mar 14;24(6):e202200703. doi: 10.1002/cbic.202200703. Epub 2023 Feb 21. Chembiochem. 2023. PMID: 36624049 Review.
-
Hyperpolarized NMR Spectroscopy: d-DNP, PHIP, and SABRE Techniques.Chem Asian J. 2018 May 23:10.1002/asia.201800551. doi: 10.1002/asia.201800551. Online ahead of print. Chem Asian J. 2018. PMID: 29790649 Free PMC article. Review.
Cited by
-
Biphasic NMR of Hyperpolarized Suspensions-Real-Time Monitoring of Solute-to-Solid Conversion to Watch Materials Grow.J Phys Chem C Nanomater Interfaces. 2023 Sep 21;127(39):19591-19598. doi: 10.1021/acs.jpcc.3c04198. eCollection 2023 Oct 5. J Phys Chem C Nanomater Interfaces. 2023. PMID: 37817917 Free PMC article.
-
Unconventional Parahydrogen-Induced Hyperpolarization Effects in Chemistry and Catalysis: From Photoreactions to Enzymes.ACS Catal. 2025 Apr 4;15(8):6386-6409. doi: 10.1021/acscatal.4c07870. eCollection 2025 Apr 18. ACS Catal. 2025. PMID: 40270879 Free PMC article. Review.
-
Cross-Polarization of Insensitive Nuclei from Water Protons for Detection of Protein-Ligand Binding.J Am Chem Soc. 2024 Sep 11;146(36):24754-24758. doi: 10.1021/jacs.4c08241. Epub 2024 Sep 3. J Am Chem Soc. 2024. PMID: 39225120 Free PMC article.
-
NMR-identification of the interaction between BRCA1 and the intrinsically disordered monomer of the Myc-associated factor X.Protein Sci. 2024 Jan;33(1):e4849. doi: 10.1002/pro.4849. Protein Sci. 2024. PMID: 38037490 Free PMC article.
References
-
- Bessa L. M., Guseva S., Camacho-Zarco A. R., Salvi N., Maurin D., Perez L. M., Botova M., Malki A., Nanao M., Jensen M. R., Ruigrok R. W. H., Blackledge M., The intrinsically disordered SARS-CoV-2 nucleoprotein in dynamic complex with its viral partner nsp3a. Sci. Adv. 8, eabm4034 (2022). - PMC - PubMed
-
- Theillet F. X., Binolfi A., Bekei B., Martorana A., Rose H. M., Stuiver M., Verzini S., Lorenz D., van Rossum M., Goldfarb D., Selenko P., Structural disorder of monomeric α-synuclein persists in mammalian cells. Nature 530, 45–50 (2016). - PubMed
-
- Wilmanns M., Gautel M., Mayans O., Activation of calcium/calmodulin regulated kinases. Cell. Mol. Biol. 46, 883–894 (2000). - PubMed
LinkOut - more resources
Full Text Sources

