Temperature-dependent hydrogen deuterium exchange shows impact of analog binding on adenosine deaminase flexibility but not embedded thermal networks
- PMID: 35933011
- PMCID: PMC9483566
- DOI: 10.1016/j.jbc.2022.102350
Temperature-dependent hydrogen deuterium exchange shows impact of analog binding on adenosine deaminase flexibility but not embedded thermal networks
Abstract
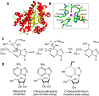
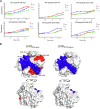
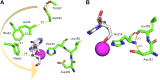
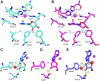
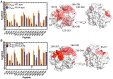
The analysis of hydrogen deuterium exchange by mass spectrometry as a function of temperature and mutation has emerged as a generic and efficient tool for the spatial resolution of protein networks that are proposed to function in the thermal activation of catalysis. In this work, we extend temperature-dependent hydrogen deuterium exchange from apo-enzyme structures to protein-ligand complexes. Using adenosine deaminase as a prototype, we compared the impacts of a substrate analog (1-deaza-adenosine) and a very tight-binding inhibitor/transition state analog (pentostatin) at single and multiple temperatures. At a single temperature, we observed different hydrogen deuterium exchange-mass spectrometry properties for the two ligands, as expected from their 106-fold differences in strength of binding. By contrast, analogous patterns for temperature-dependent hydrogen deuterium exchange mass spectrometry emerge in the presence of both 1-deaza-adenosine and pentostatin, indicating similar impacts of either ligand on the enthalpic barriers for local protein unfolding. We extended temperature-dependent hydrogen deuterium exchange to a function-altering mutant of adenosine deaminase in the presence of pentostatin and revealed a protein thermal network that is highly similar to that previously reported for the apo-enzyme (Gao et al., 2020, JACS 142, 19936-19949). Finally, we discuss the differential impacts of pentostatin binding on overall protein flexibility versus site-specific thermal transfer pathways in the context of models for substrate-induced changes to a distributed protein conformational landscape that act in synergy with embedded protein thermal networks to achieve efficient catalysis.
Keywords: adenosine deaminase; conformational landscape; protein flexibility; temperature-dependent hydrogen deuterium exchange (TDHDX-MS); thermal activation.
Copyright © 2022 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare no conflict of interest.
Figures



Similar articles
-
Hydrogen-Deuterium Exchange within Adenosine Deaminase, a TIM Barrel Hydrolase, Identifies Networks for Thermal Activation of Catalysis.J Am Chem Soc. 2020 Nov 25;142(47):19936-19949. doi: 10.1021/jacs.0c07866. Epub 2020 Nov 12. J Am Chem Soc. 2020. PMID: 33181018 Free PMC article.
-
Dynamical activation of function in metalloenzymes.FEBS Lett. 2023 Jan;597(1):79-91. doi: 10.1002/1873-3468.14515. Epub 2022 Oct 27. FEBS Lett. 2023. PMID: 36239559 Free PMC article. Review.
-
Measuring the hydrogen/deuterium exchange of proteins at high spatial resolution by mass spectrometry: overcoming gas-phase hydrogen/deuterium scrambling.Acc Chem Res. 2014 Oct 21;47(10):3018-27. doi: 10.1021/ar500194w. Epub 2014 Aug 29. Acc Chem Res. 2014. PMID: 25171396 Review.
-
Temperature-induced conformational change at the catalytic site of Sulfolobus solfataricus alcohol dehydrogenase highlighted by Asn249Tyr substitution. A hydrogen/deuterium exchange, kinetic, and fluorescence quenching study.Biochemistry. 2005 Aug 23;44(33):11040-8. doi: 10.1021/bi050469c. Biochemistry. 2005. PMID: 16101287
-
Thermal denaturation of Escherichia coli thioredoxin studied by hydrogen/deuterium exchange and electrospray ionization mass spectrometry: monitoring a two-state protein unfolding transition.Biochemistry. 1999 Jan 19;38(3):1136-43. doi: 10.1021/bi981938w. Biochemistry. 1999. PMID: 9894011
Cited by
-
A Foundational Shift in Models for Enzyme Function.J Am Chem Soc. 2025 May 7;147(18):14884-14904. doi: 10.1021/jacs.5c02388. Epub 2025 Apr 25. J Am Chem Soc. 2025. PMID: 40277147 Free PMC article. Review.
-
Identification of Scaffold Specific Energy Transfer Networks in the Enthalpic Activation of Orotidine 5'-Monophosphate Decarboxylase.bioRxiv [Preprint]. 2025 Jan 29:2025.01.29.635545. doi: 10.1101/2025.01.29.635545. bioRxiv. 2025. PMID: 39975186 Free PMC article. Preprint.
-
Identification of the Thermal Activation Network in Human 15-Lipoxygenase-2: Divergence from Plant Orthologs and Its Relationship to Hydrogen Tunneling Activation Barriers.ACS Catal. 2024 Mar 28;14(7):5444-5457. doi: 10.1021/acscatal.4c00439. eCollection 2024 Apr 5. ACS Catal. 2024. PMID: 38601784 Free PMC article.
-
Examining DNA Structures with In-droplet Hydrogen/Deuterium Exchange Mass Spectrometry.Int J Mass Spectrom. 2024 May;499:117231. doi: 10.1016/j.ijms.2024.117231. Epub 2024 Mar 7. Int J Mass Spectrom. 2024. PMID: 38854816 Free PMC article.
-
Temporal Resolution of Activity-Related Solvation Dynamics in the TIM Barrel Enzyme Murine Adenosine Deaminase.ACS Catal. 2024 Apr 5;14(7):4554-4567. doi: 10.1021/acscatal.3c02687. Epub 2024 Mar 12. ACS Catal. 2024. PMID: 39099600 Free PMC article.
References
-
- Lipscomb W.N. Acceleration of reactions by enzymes. Acc. Chem. Res. 1982;15:232–238.
-
- Kraut J. How do enzymes work? Science. 1988;242:533–540. - PubMed
-
- Radzicka A., Wolfenden R. A proficient enzyme. Science. 1995;267:90–93. - PubMed
-
- Miller B.G., Wolfenden R. Catalytic proficiency: the unusual case of OMP decarboxylase. Annu. Rev. Biochem. 2002;71:847–885. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

