Intra-species diversity of Clostridium perfringens: A diverse genetic repertoire reveals its pathogenic potential
- PMID: 35935202
- PMCID: PMC9354469
- DOI: 10.3389/fmicb.2022.952081
Intra-species diversity of Clostridium perfringens: A diverse genetic repertoire reveals its pathogenic potential
Abstract
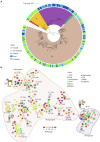
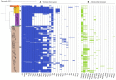
Clostridium perfringens is the causative agent of many enterotoxic diseases in humans and animals, and it is present in diverse environments (soil, food, sewage, and water). Multilocus Sequence Typing (MLST) and Whole Genome Sequencing (WGS) have provided a general approach about genetic diversity of C. perfringens; however, those studies are limited to specific locations and often include a reduced number of genomes. In this study, 372 C. perfringens genomes from multiple locations and sources were used to assess the genetic diversity and phylogenetic relatedness of this pathogen. In silico MLST was used for typing the isolates, and the resulting sequence types (ST) were assigned to clonal complexes (CC) based on allelic profiles that differ from its founder by up to double-locus variants. A pangenome analysis was conducted, and a core genome-based phylogenetic tree was created to define phylogenetic groups. Additionally, key virulence factors, toxinotypes, and antibiotic resistance genes were identified using ABRicate against Virulence Factor Database (VFDB), TOXiper, and Resfinder, respectively. The majority of the C. perfringens genomes found in publicly available databases were derived from food (n = 85) and bird (n = 85) isolates. A total of 195 STs, some of them shared between sources such as food and human, horses and dogs, and environment and birds, were grouped in 25 CC and distributed along five phylogenetic groups. Fifty-three percent of the genomes were allocated to toxinotype A, followed by F (32%) and G (7%). The most frequently found virulence factors based on > 70% coverage and 99.95% identity were plc (100%), nanH (99%), ccp (99%), and colA (98%), which encode an alpha-toxin, a sialidase, an alpha-clostripain, and a collagenase, respectively, while tetA (39.5%) and tetB (36.2%), which mediate tetracycline resistance determinants, were the most common antibiotic resistance genes detected. The analyses conducted here showed a better view of the presence of this pathogen across several host species. They also confirm that the genetic diversity of C. perfringens is based on a large number of virulence factors that vary among phylogroups, and antibiotic resistance markers, especially to tetracyclines, aminoglycosides, and macrolides. Those characteristics highlight the importance of C. perfringens as a one of the most common causes of foodborne illness.
Keywords: Clostridium perfringens; genomic epidemiology; intra-species diversity; multilocus sequence typing; toxinotypes.
Copyright © 2022 Camargo, Guerrero-Araya, Castañeda, Vega, Cardenas-Alvarez, Rodríguez, Paredes-Sabja, Ramírez and Muñoz.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Abdel-Glil M. Y., Thomas P., Linde J., Busch A., Wieler L. H., Neubauer H., et al. . (2021). Comparative in silico genome analysis of Clostridium perfringens unravels stable phylogroups with different genome characteristics and pathogenic potential. Sci. Rep. 11:6756. doi: 10.1038/s41598-021-86148-8, PMID: - DOI - PMC - PubMed
-
- Al-Shukri M. S., Hmood A. M., Al-Charrakh A. H. (2021). Sequencing of Clostridium perfringens toxin genes (cpa, etx, iap) from Iraqi hospitals and detection by PCR of the genes encoding resistance to metronidazole, tetracycline, and clindamycin. Indian J. Med. Microbiol. 39, 289–294. doi: 10.1016/j.ijmmb.2021.03.017, PMID: - DOI - PubMed
-
- Awad M. M., Bryant A. E., Stevens D. L., Rood J. I. (1995). Virulence studies on chromosomal α-toxin and Θ-toxin mutants constructed by allelic exchange provide genetic evidence for the essential role of α-toxin in Clostridium perfringens-mediated gas gangrene. Mol. Microbiol. 15, 191–202. doi: 10.1111/j.1365-2958.1995.tb02234.x, PMID: - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

