Transcriptomic analysis of genes related to alkaloid biosynthesis and the regulation mechanism under precursor and methyl jasmonate treatment in Dendrobium officinale
- PMID: 35937364
- PMCID: PMC9355482
- DOI: 10.3389/fpls.2022.941231
Transcriptomic analysis of genes related to alkaloid biosynthesis and the regulation mechanism under precursor and methyl jasmonate treatment in Dendrobium officinale
Abstract
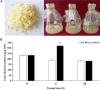
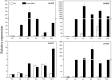
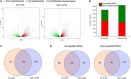
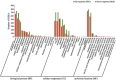
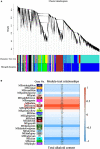
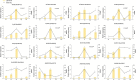
Dendrobium officinale is both a traditional herbal medicine and a plant of high ornamental and medicinal value. Alkaloids, especially terpenoid indole alkaloids (TIAs), with pharmacological activities are present in the tissues of D. officinale. A number of genes involved in alkaloid biosynthetic pathways have been identified. However, the regulatory mechanisms underlying the precursor and methyl jasmonate (MeJA)-induced accumulation of alkaloids in D. officinale are poorly understood. In this study, we collected D. officinale protocorm-like bodies (PLBs) and treated them with TIA precursors (tryptophan and secologanin) and MeJA for 0 (T0), 4 (T4) and 24 h (T24); we also established control samples (C4 and C24). Then, we measured the total alkaloid content of the PLBs and performed transcriptome sequencing using the Illumina HiSeq 2,500 system. The total alkaloid content increased significantly after 4 h of treatment. Go and KEGG analysis suggested that genes from the TIA, isoquinoline alkaloid, tropane alkaloid and jasmonate (JA) biosynthetic pathways were significantly enriched. Weighted gene coexpression network analysis (WGCNA) uncovered brown module related to alkaloid content. Six and seven genes related to alkaloid and JA bisosynthetic pathways, respectively, might encode the key enzymes involved in alkaloid biosynthesis of D. officinale. Moreover, 13 transcription factors (TFs), which mostly belong to AP2/ERF, WRKY, and MYB gene families, were predicted to regulate alkaloid biosynthesis. Our data provide insight for studying the regulatory mechanism underlying TIA precursor and MeJA-induced accumulation of three types of alkaloids in D. officinale.
Keywords: Dendrobium officinale; alkaloid biosynthesis; methyl jasmonate; precursors; transcriptome.
Copyright © 2022 Jiao, Wei, Fan, Song, Wang, Cai and Jin.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The handling editor, YC, declared a past co-authorship with the authors, YC and CS.
Figures




Similar articles
-
Metabolic Profiling of Dendrobium officinale in Response to Precursors and Methyl Jasmonate.Int J Mol Sci. 2018 Mar 3;19(3):728. doi: 10.3390/ijms19030728. Int J Mol Sci. 2018. PMID: 29510516 Free PMC article.
-
Comparative transcriptomic analysis reveal the regulation mechanism underlying MeJA-induced accumulation of alkaloids in Dendrobium officinale.J Plant Res. 2019 May;132(3):419-429. doi: 10.1007/s10265-019-01099-6. Epub 2019 Mar 22. J Plant Res. 2019. PMID: 30903398
-
Putative genes in alkaloid biosynthesis identified in Dendrobium officinale by correlating the contents of major bioactive metabolites with genes expression between Protocorm-like bodies and leaves.BMC Genomics. 2021 Jul 29;22(1):579. doi: 10.1186/s12864-021-07887-6. BMC Genomics. 2021. PMID: 34325653 Free PMC article.
-
Transcription factors in alkaloid biosynthesis.Int Rev Cell Mol Biol. 2013;305:339-82. doi: 10.1016/B978-0-12-407695-2.00008-1. Int Rev Cell Mol Biol. 2013. PMID: 23890386 Review.
-
Natural Composition and Biosynthetic Pathways of Alkaloids in Medicinal Dendrobium Species.Front Plant Sci. 2022 May 6;13:850949. doi: 10.3389/fpls.2022.850949. eCollection 2022. Front Plant Sci. 2022. PMID: 35599884 Free PMC article. Review.
Cited by
-
Metabolic and Transcriptomic Profile Revealing the Differential Accumulating Mechanism in Different Parts of Dendrobium nobile.Int J Mol Sci. 2024 May 14;25(10):5356. doi: 10.3390/ijms25105356. Int J Mol Sci. 2024. PMID: 38791394 Free PMC article.
-
In-depth analysis of large-scale screening of WRKY members based on genome-wide identification.Front Genet. 2023 Jan 9;13:1104968. doi: 10.3389/fgene.2022.1104968. eCollection 2022. Front Genet. 2023. PMID: 36699467 Free PMC article.
-
Tomato Defenses Under Stress: The Impact of Salinity on Direct Defenses Against Insect Herbivores.Plant Cell Environ. 2025 May;48(5):3647-3659. doi: 10.1111/pce.15353. Epub 2025 Jan 13. Plant Cell Environ. 2025. PMID: 39806825 Free PMC article.
-
Genome-wide identification and expression analysis of TIFY genes under MeJA, cold and PEG-induced drought stress treatment in Dendrobium huoshanense.Physiol Mol Biol Plants. 2024 Apr;30(4):527-542. doi: 10.1007/s12298-024-01442-9. Epub 2024 Mar 25. Physiol Mol Biol Plants. 2024. PMID: 38737319 Free PMC article.
-
Identification of Dof transcription factors in Dendrobium huoshanense and expression pattern under abiotic stresses.Front Genet. 2024 Apr 22;15:1394790. doi: 10.3389/fgene.2024.1394790. eCollection 2024. Front Genet. 2024. PMID: 38711915 Free PMC article.
References
-
- Chen C., Zhu H., Kang J., Warusawitharana H. K., Chen S., Wang K., et al. (2022). Comparative transcriptome and phytochemical analysis provides insight into triterpene saponin biosynthesis in seeds and flowers of the tea plant (Camellia sinensis). Metabolites 12:204. 10.3390/metabo12030204 - DOI - PMC - PubMed
-
- Fan H., Wu Q., Wang X., Wu L., Cai Y., Lin Y. (2016). Molecular cloning and expression of 1-deoxy-d-xylulose-5-phosphate synthase and 1-deoxy-d-xylulose-5-phosphate reductoisomerase in Dendrobium officinale. Plant Cell Tiss. Organ Cult. 125 381–385. 10.1007/s11240-016-0945-1 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

