Exploring the role of antimicrobials in the selective growth of purple phototrophic bacteria through genome mining and agar spot assays
- PMID: 35938312
- PMCID: PMC9804395
- DOI: 10.1111/lam.13795
Exploring the role of antimicrobials in the selective growth of purple phototrophic bacteria through genome mining and agar spot assays
Abstract
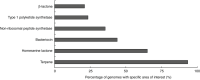
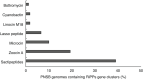
Purple non-sulphur bacteria (PNSB) are an emerging group of microbes attractive for applied microbiology applications such as wastewater treatment, plant biostimulants, microbial protein, polyhydroxyalkanoates and H2 production. These photoorganoheterotrophic microbes have the unique ability to grow selectively on organic carbon in anaerobic photobioreactors. This so-called selectivity implies that the microbial community will have a low diversity and a high abundance of a particular PNSB species. Recently, it has been shown that certain PNSB strains can produce antimicrobials, yet it remains unclear whether these contribute to competitive inhibition. This research aimed to understand which type of antimicrobial PNSB produce and identify whether these compounds contribute to their selective growth. Mining 166 publicly-available PNSB genomes using the computational tool BAGEL showed that 59% contained antimicrobial encoding regions, more specifically biosynthetic clusters of bacteriocins and non-ribosomal peptide synthetases. Inter- and intra-species inhibition was observed in agar spot assays for Rhodobacter blasticus EBR2 and Rhodopseudomonas palustris EBE1 with inhibition zones of, respectively, 5.1 and 1.5-5.7 mm. Peptidomic analysis detected a peptide fragment in the supernatant (SVLQLLR) that had a 100% percentage identity match with a known non-ribosomal peptide synthetase with antimicrobial activity.
Keywords: alternative protein; animal feed; antibiotics; antimicrobial peptide; bacteriocin; probiotic; purple phototrophic bacteria.
© 2022 The Authors. Letters in Applied Microbiology published by John Wiley & Sons Ltd on behalf of Society for Applied Microbiology.
Conflict of interest statement
No conflict of interest declared.
Figures
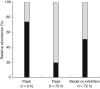
 ) at time (t) 0 h, 72 h and through the growth model. The PNSB abundancies in the growth model were calculated using the equation N = N0*eμmax*
(tfinal−tlag) without considering competitive inhibition. The abundance was corrected for 16S gene copies per genome.
) at time (t) 0 h, 72 h and through the growth model. The PNSB abundancies in the growth model were calculated using the equation N = N0*eμmax*
(tfinal−tlag) without considering competitive inhibition. The abundance was corrected for 16S gene copies per genome.Similar articles
-
Volatile fatty acids impacting phototrophic growth kinetics of purple bacteria: Paving the way for protein production on fermented wastewater.Water Res. 2019 Apr 1;152:138-147. doi: 10.1016/j.watres.2018.12.025. Epub 2018 Dec 27. Water Res. 2019. PMID: 30665160
-
Anoxygenic phototrophic purple non-sulfur bacteria: tool for bioremediation of hazardous environmental pollutants.World J Microbiol Biotechnol. 2023 Aug 18;39(10):283. doi: 10.1007/s11274-023-03729-7. World J Microbiol Biotechnol. 2023. PMID: 37594588 Free PMC article. Review.
-
Operational Strategies to Selectively Produce Purple Bacteria for Microbial Protein in Raceway Reactors.Environ Sci Technol. 2021 Jun 15;55(12):8278-8286. doi: 10.1021/acs.est.0c08204. Epub 2021 Jun 4. Environ Sci Technol. 2021. PMID: 34085818
-
Dehazing redox homeostasis to foster purple bacteria biotechnology.Trends Biotechnol. 2023 Jan;41(1):106-119. doi: 10.1016/j.tibtech.2022.06.010. Epub 2022 Jul 14. Trends Biotechnol. 2023. PMID: 35843758 Review.
-
Cocultivating aerobic heterotrophs and purple bacteria for microbial protein in sequential photo- and chemotrophic reactors.Bioresour Technol. 2021 Jan;319:124192. doi: 10.1016/j.biortech.2020.124192. Epub 2020 Sep 30. Bioresour Technol. 2021. PMID: 33039841
References
-
- Abrudan, M.I. , Brown, S. and Rozen, D.E. (2012) Killing as means of promoting biodiversity. Biochem Soc Trans 40, 1512–1516. - PubMed
-
- Agler, M.T. , Wrenn, B.A. , Zinder, S.H. and Angenent, L.T. (2011) Waste to bioproduct conversion with undefined mixed cultures: the carboxylate platform. Trends Biotechnol 29, 70–78. - PubMed
-
- Alloul, A. , Cerruti, M. , Adamczyk, D. , Weissbrodt, D.G. and Vlaeminck, S.E. (2021) Operational strategies to selectively produce purple bacteria for microbial protein in raceway reactors. Environ Sci Technol 55, 8278–8286. - PubMed
-
- Alloul, A. , Muys, M. , Hertoghs, N. , Kerckhof, F.‐M. and Vlaeminck, S.E. (2021) Cocultivating aerobic heterotrophs and purple bacteria for microbial protein in sequential photo‐and chemotrophic reactors. Bioresour Technol 319, 124192. - PubMed

