TMEM41B and VMP1 modulate cellular lipid and energy metabolism for facilitating dengue virus infection
- PMID: 35939522
- PMCID: PMC9387935
- DOI: 10.1371/journal.ppat.1010763
TMEM41B and VMP1 modulate cellular lipid and energy metabolism for facilitating dengue virus infection
Abstract
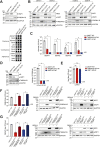
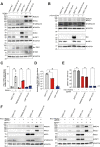
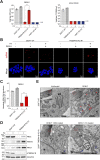
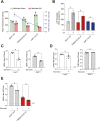
Transmembrane Protein 41B (TMEM41B) and Vacuole Membrane Protein 1 (VMP1) are two ER-associated lipid scramblases that play a role in autophagosome formation and cellular lipid metabolism. TMEM41B is also a recently validated host factor required by flaviviruses and coronaviruses. However, the exact underlying mechanism of TMEM41B in promoting viral infections remains an open question. Here, we validated that both TMEM41B and VMP1 are essential host dependency factors for all four serotypes of dengue virus (DENV) and human coronavirus OC43 (HCoV-OC43), but not chikungunya virus (CHIKV). While HCoV-OC43 failed to replicate entirely in both TMEM41B- and VMP1-deficient cells, we detected diminished levels of DENV infections in these cell lines, which were accompanied by upregulation of the innate immune dsRNA sensors, RIG-I and MDA5. Nonetheless, this upregulation did not correspondingly induce the downstream effector TBK1 activation and Interferon-beta expression. Despite low levels of DENV replication, classical DENV replication organelles were undetectable in the infected TMEM41B-deficient cells, suggesting that the upregulation of the dsRNA sensors is likely a consequence of aberrant viral replication rather than a causal factor for reduced DENV infection. Intriguingly, we uncovered that the inhibitory effect of TMEM41B deficiency on DENV replication, but not HCoV-OC43, can be partially reversed using exogenous fatty acid supplements. In contrast, VMP1 deficiency cannot be rescued using the metabolite treatment. In line with the observed phenotypes, we found that both TMEM41B- and VMP1-deficient cells harbor higher levels of compromised mitochondria, especially in VMP1 deficiency which results in severe dysregulations of mitochondrial beta-oxidation. Using a metabolomic profiling approach, we revealed distinctive global dysregulations of the cellular metabolome, particularly lipidome, in TMEM41B- and VMP1-deficient cells. Our findings highlight a central role for TMEM41B and VMP1 in modulating multiple cellular pathways, including lipid mobilization, mitochondrial beta-oxidation, and global metabolic regulations, to facilitate the replication of flaviviruses and coronaviruses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
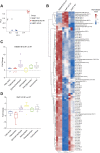

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

