Geographic patterns of koala retrovirus genetic diversity, endogenization, and subtype distributions
- PMID: 35943984
- PMCID: PMC9388103
- DOI: 10.1073/pnas.2122680119
Geographic patterns of koala retrovirus genetic diversity, endogenization, and subtype distributions
Erratum in
-
Correction for Blyton et al., Geographic patterns of koala retrovirus genetic diversity, endogenization, and subtype distributions.Proc Natl Acad Sci U S A. 2023 Jan 3;120(1):e2220541120. doi: 10.1073/pnas.2220541120. Epub 2022 Dec 29. Proc Natl Acad Sci U S A. 2023. PMID: 36580601 Free PMC article. No abstract available.
Abstract
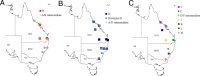
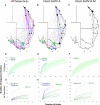
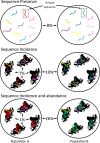
Koala retrovirus (KoRV) subtype A (KoRV-A) is currently in transition from exogenous virus to endogenous viral element, providing an ideal system to elucidate retroviral-host coevolution. We characterized KoRV geography using fecal DNA from 192 samples across 20 populations throughout the koala's range. We reveal an abrupt change in KoRV genetics and incidence at the Victoria/New South Wales state border. In northern koalas, pol gene copies were ubiquitously present at above five per cell, consistent with endogenous KoRV. In southern koalas, pol copies were detected in only 25.8% of koalas and always at copy numbers below one, while the env gene was detected in all animals and in a majority at copy numbers above one per cell. These results suggest that southern koalas carry partial endogenous KoRV-like sequences. Deep sequencing of the env hypervariable region revealed three putatively endogenous KoRV-A sequences in northern koalas and a single, distinct sequence present in all southern koalas. Among northern populations, env sequence diversity decreased with distance from the equator, suggesting infectious KoRV-A invaded the koala genome in northern Australia and then spread south. The exogenous KoRV subtypes (B to K), two novel subtypes, and intermediate subtypes were detected in all northern koala populations but were strikingly absent from all southern animals tested. Apart from KoRV subtype D, these exogenous subtypes were generally locally prevalent but geographically restricted, producing KoRV genetic differentiation among northern populations. This suggests that sporadic evolution and local transmission of the exogenous subtypes have occurred within northern Australia, but this has not extended into animals within southern Australia.
Keywords: endogenization; evolution; genetic diversity; geography; koala retrovirus.
Conflict of interest statement
The authors declare no competing interest.
Figures




Similar articles
-
Phylogenetic and geographical analysis of a retrovirus during the early stages of endogenous adaptation and exogenous spread in a new host.Mol Ecol. 2021 Jun;30(11):2626-2640. doi: 10.1111/mec.15735. Epub 2020 Dec 30. Mol Ecol. 2021. PMID: 33219558 Free PMC article.
-
Phylogenetic Diversity of Koala Retrovirus within a Wild Koala Population.J Virol. 2017 Jan 18;91(3):e01820-16. doi: 10.1128/JVI.01820-16. Print 2017 Feb 1. J Virol. 2017. PMID: 27881645 Free PMC article.
-
Molecular Dynamics and Mode of Transmission of Koala Retrovirus as It Invades and Spreads through a Wild Queensland Koala Population.J Virol. 2018 Mar 1;92(5):e01871-17. doi: 10.1128/JVI.01871-17. Epub 2017 Dec 13. J Virol. 2018. PMID: 29237837 Free PMC article.
-
Koala Retroviruses: Evolution and Disease Dynamics.Annu Rev Virol. 2015 Nov;2(1):119-34. doi: 10.1146/annurev-virology-100114-055056. Annu Rev Virol. 2015. PMID: 26958909 Review.
-
Koala retrovirus epidemiology, transmission mode, pathogenesis, and host immune response in koalas (Phascolarctos cinereus): a review.Arch Virol. 2020 Nov;165(11):2409-2417. doi: 10.1007/s00705-020-04770-9. Epub 2020 Aug 8. Arch Virol. 2020. PMID: 32770481 Free PMC article. Review.
Cited by
-
Diversity and transmission of koala retrovirus: a case study in three captive koala populations.Sci Rep. 2022 Sep 22;12(1):15787. doi: 10.1038/s41598-022-18939-6. Sci Rep. 2022. PMID: 36138048 Free PMC article.
-
Elevated oncogene expressions in koala infected with multiple koala retrovirus subtypes: a preliminary study.Virus Genes. 2025 Jun 26. doi: 10.1007/s11262-025-02169-9. Online ahead of print. Virus Genes. 2025. PMID: 40569497
-
FeLIX is a restriction factor for mammalian retrovirus infection.J Virol. 2024 Apr 16;98(4):e0177123. doi: 10.1128/jvi.01771-23. Epub 2024 Mar 5. J Virol. 2024. PMID: 38440982 Free PMC article.
-
Koala retrovirus load and non-A subtypes are associated with secondary disease among wild northern koalas.PLoS Pathog. 2022 May 19;18(5):e1010513. doi: 10.1371/journal.ppat.1010513. eCollection 2022 May. PLoS Pathog. 2022. PMID: 35588407 Free PMC article.
-
Genome Reference Assembly for Bottlenecked Southern Australian Koalas.Genome Biol Evol. 2023 Jan 4;15(1):evac176. doi: 10.1093/gbe/evac176. Genome Biol Evol. 2023. PMID: 36542479 Free PMC article.
References
-
- Bock M., Stoye J. P., Endogenous retroviruses and the human germline. Curr. Opin. Genet. Dev. 10, 651–655 (2000). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

