Massively Parallel Selection of NanoCluster Beacons
- PMID: 35945159
- PMCID: PMC9588665
- DOI: 10.1002/adma.202204957
Massively Parallel Selection of NanoCluster Beacons
Abstract
NanoCluster Beacons (NCBs) are multicolor silver nanocluster probes whose fluorescence can be activated or tuned by a proximal DNA strand called the activator. While a single-nucleotide difference in a pair of activators can lead to drastically different activation outcomes, termed polar opposite twins (POTs), it is difficult to discover new POT-NCBs using the conventional low-throughput characterization approaches. Here, a high-throughput selection method is reported that takes advantage of repurposed next-generation-sequencing chips to screen the activation fluorescence of ≈40 000 activator sequences. It is found that the nucleobases at positions 7-12 of the 18-nucleotide-long activator are critical to creating bright NCBs and positions 4-6 and 2-4 are hotspots to generate yellow-orange and red POTs, respectively. Based on these findings, a "zipper-bag" model is proposed that can explain how these hotspots facilitate the formation of distinct silver cluster chromophores and alter their chemical yields. Combining high-throughput screening with machine-learning algorithms, a pipeline is established to design bright and multicolor NCBs in silico.
Keywords: NanoCluster Beacons; fluorescent nanomaterials; high-throughput screening; next-generation sequencing; silver nanoclusters.
© 2022 Wiley-VCH GmbH.
Conflict of interest statement
Competing interests
The authors declare no competing interests.
Figures



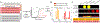
References
-
- Yeh HC, Sharma J, Han JJ, Martinez JS, Werner JH, Nano Letters 2010, 10, 3106. - PubMed
-
- Yeh H-C, Sharma J, Han JJ, Martinez JS, Werner JH, IEEE Nanotechnology Magazine 2011, 5, 28.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

