Tight junction formation by a claudin mutant lacking the COOH-terminal PDZ domain-binding motif
- PMID: 35945631
- PMCID: PMC9805020
- DOI: 10.1111/nyas.14881
Tight junction formation by a claudin mutant lacking the COOH-terminal PDZ domain-binding motif
Abstract
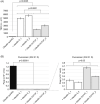
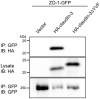
Claudin-based tight junctions (TJs) are formed at the most apical part of cell-cell contacts in epithelial cells. Previous studies suggest that scaffolding proteins ZO-1 and ZO-2 (ZO proteins) determine the location of TJs by interacting with claudins, but this idea is not conclusive. To address the role of the ZO proteins binding to claudins at TJs, a COOH-terminal PDZ domain binding motif-deleted claudin-3 mutant, which lacks the ZO protein binding, was stably expressed in claudin-deficient MDCK cells. The COOH-terminus-deleted claudin-3 was localized at the apicolateral region similar to full-length claudin-3. Consistently, freeze-fracture electron microscopy revealed that the COOH-terminus-deleted claudin-3-expressing cells reconstituted belts of TJs at the most apical region of the lateral membrane and restored functional epithelial barriers. These results suggest that the interaction of claudins with ZO proteins is not a prerequisite for TJ formation at the most apical part of cell-cell contacts.
Keywords: PDZ domain-binding motif; ZO-1; claudin; tight junction.
© 2022 The Authors. Annals of the New York Academy of Sciences published by Wiley Periodicals LLC on behalf of New York Academy of Sciences.
Conflict of interest statement
The authors have no competing interests to declare.
Figures




References
-
- Otani, T. , & Furuse, M. (2020). Tight junction structure and function revisited. Trends in Cell Biology, 30, 805–817. - PubMed
-
- Zihni, C. , Mills, C. , Matter, K. , & Balda, M. S. (2016). Tight junctions: From simple barriers to multifunctional molecular gates. Nature Reviews Molecular Cell Biology, 17, 564–580. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

