Co-option of the piRNA pathway to regulate neural crest specification
- PMID: 35947657
- PMCID: PMC9365273
- DOI: 10.1126/sciadv.abn1441
Co-option of the piRNA pathway to regulate neural crest specification
Abstract
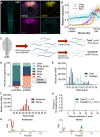
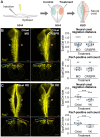
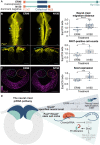
Across Metazoa, Piwi proteins play a critical role in protecting the germline genome through piRNA-mediated repression of transposable elements. In vertebrates, activity of Piwi proteins and the piRNA pathway was thought to be gonad specific. Our results reveal the expression of Piwil1 in a vertebrate somatic cell type, the neural crest. Piwil1 is expressed at low levels throughout the chicken neural tube, peaking in neural crest cells just before the specification event that enables epithelial-to-mesenchymal transition (EMT) and migration into the periphery. Loss of Piwil1 impedes neural crest specification and emigration. Small RNA sequencing reveals somatic piRNAs with sequence signatures of an active ping-pong loop. RNA-seq and functional experiments identify the transposon-derived gene ERNI as Piwil1's target in the neural crest. ERNI, in turn, suppresses Sox2 to precisely control the timing of neural crest specification and EMT. Our data provide mechanistic insight into a novel function of the piRNA pathway as a regulator of somatic development in a vertebrate species.
Figures

References
-
- Cenik E. S., Zamore P. D., Argonaute proteins. Curr. Biol. 21, R446–R449 (2011). - PubMed
-
- Aravin A. A., Sachidanandam R., Girard A., Fejes-Toth K., Hannon G. J., Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007). - PubMed
-
- Brennecke J., Aravin A. A., Stark A., Dus M., Kellis M., Sachidanandam R., Hannon G. J., Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007). - PubMed
-
- Vagin V. V., Sigova A., Li C., Seitz H., Gvozdev V., Zamore P. D., A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006). - PubMed
-
- Aravin A. A., Hannon G. J., Brennecke J., The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 (2007). - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

