Network topology-directed design of molecular CPU for cell-like dynamic information processing
- PMID: 35947658
- PMCID: PMC9365278
- DOI: 10.1126/sciadv.abq0917
Network topology-directed design of molecular CPU for cell-like dynamic information processing
Abstract
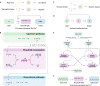
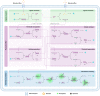
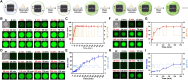
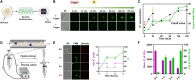
Natural cells (NCs) can automatically and continuously respond to fluctuant external information and distinguish meaningful stimuli from weak noise depending on their powerful genetic and protein networks. We herein report a network topology-directed design of dynamic molecular processing system (DMPS) as a molecular central processing unit that powers an artificial cell (AC) able to process fluctuant information in its immediate environment similar to NCs. By constructing a mixed cell community, ACs and NCs have synchronous response to fluctuant extracellular stimuli under physiological condition and in a blood vessel-mimic circulation system. We also show that fluctuant bioinformation released by NCs can be received and processed by ACs. The molecular design of DMPS-powered AC is expected to allow a profound understanding of biological systems, advance the construction of intelligent molecular systems, and promote more elegant bioengineering applications.
Figures



Similar articles
-
An information theoretic approach to constructing robust Boolean gene regulatory networks.IEEE/ACM Trans Comput Biol Bioinform. 2012 Jan-Feb;9(1):52-65. doi: 10.1109/TCBB.2011.61. Epub 2011 Mar 22. IEEE/ACM Trans Comput Biol Bioinform. 2012. PMID: 21464507
-
Simplified activity cliff network representations with high interpretability and immediate access to SAR information.J Comput Aided Mol Des. 2020 Sep;34(9):943-952. doi: 10.1007/s10822-020-00319-9. Epub 2020 Jun 5. J Comput Aided Mol Des. 2020. PMID: 32500478 Free PMC article.
-
A cascade reaction network mimicking the basic functional steps of adaptive immune response.Nat Chem. 2015 Oct;7(10):835-41. doi: 10.1038/nchem.2325. Epub 2015 Aug 17. Nat Chem. 2015. PMID: 26391084 Free PMC article.
-
Digital and analog chemical evolution.Acc Chem Res. 2012 Dec 18;45(12):2189-99. doi: 10.1021/ar300214w. Epub 2012 Oct 25. Acc Chem Res. 2012. PMID: 23098254 Review.
-
The rise of intelligent matter.Nature. 2021 Jun;594(7863):345-355. doi: 10.1038/s41586-021-03453-y. Epub 2021 Jun 16. Nature. 2021. PMID: 34135518 Review.
Cited by
-
Robust Covalent Aptamer Strategy Enables Sensitive Detection and Enhanced Inhibition of SARS-CoV-2 Proteins.ACS Cent Sci. 2023 Jan 2;9(1):72-83. doi: 10.1021/acscentsci.2c01263. eCollection 2023 Jan 25. ACS Cent Sci. 2023. PMID: 36712483 Free PMC article.
-
DNA as a universal chemical substrate for computing and data storage.Nat Rev Chem. 2024 Mar;8(3):179-194. doi: 10.1038/s41570-024-00576-4. Epub 2024 Feb 9. Nat Rev Chem. 2024. PMID: 38337008 Review.
References
-
- Omidvar M., Zdarta J., Sigurdardottir S. B., Pinelo M., Mimicking natural strategies to create multi-environment enzymatic reactors: From natural cell compartments to artificial polyelectrolyte reactors. Biotechnol. Adv. 54, 107798 (2021). - PubMed
-
- Wang X., Tian L., Ren Y., Zhao Z., Du H., Zhang Z., Drinkwater B. W., Mann S., Han X., Chemical information exchange in organized protocells and natural cell assemblies with controllable spatial positions. Small 16, e1906394 (2020). - PubMed
-
- Rateitschak K., Wolkenhauer O., Thresholds in transient dynamics of signal transduction pathways. J. Theor. Biol. 264, 334–346 (2010). - PubMed
LinkOut - more resources
Full Text Sources

