A Comprehensive Transcriptomic Analysis of Arsenic-Induced Bladder Carcinogenesis
- PMID: 35954277
- PMCID: PMC9367831
- DOI: 10.3390/cells11152435
A Comprehensive Transcriptomic Analysis of Arsenic-Induced Bladder Carcinogenesis
Abstract
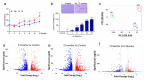
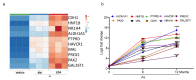
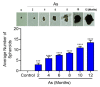
Arsenic (sodium arsenite: NaAsO2) is a potent carcinogen and a known risk factor for the onset of bladder carcinogenesis. The molecular mechanisms that govern arsenic-induced bladder carcinogenesis remain unclear. We used a physiological concentration of NaAsO2 (250 nM: 33 µg/L) for the malignant transformation of normal bladder epithelial cells (TRT-HU1), exposed for over 12 months. The increased proliferation and colony-forming abilities of arsenic-exposed cells were seen after arsenic exposure from 4 months onwards. Differential gene expression (DEG) analysis revealed that a total of 1558 and 1943 (padj < 0.05) genes were deregulated in 6-month and 12-month arsenic-exposed TRT-HU1 cells. The gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis revealed that cell proliferation and survival pathways, such as the MAPK, PI3K/AKT, and Hippo signaling pathways, were significantly altered. Pathway analysis revealed that the enrichment of stem cell activators such as ALDH1A1, HNF1b, MAL, NR1H4, and CDH1 (p < 0.001) was significantly induced during the transformation compared to respective vehicle controls. Further, these results were validated by qPCR analysis, which corroborated the transcriptomic analysis. Overall, the results suggested that stem cell activators may play a significant role in facilitating the arsenic-exposed cells to gain a survival advantage, enabling the healthy epithelial cells to reprogram into a cancer stem cell phenotype, leading to malignant transformation.
Keywords: KEGG and GO enrichment; arsenic; bladder cancer; differential gene expression; stem cells.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
-
- Choudhury M.I.M., Shabnam N., Ahsan T., Ahsan S.M.A., Kabir M.S., Khan R.M., Miah M.A., Uddin M.K., Liton M.A.R., Chakraborty S. Cutaneous Malignancy due to Arsenicosis in Bangladesh: 12-Year Study in Tertiary Level Hospital. Biomed Res. Int. 2018;2018:4678362. doi: 10.1155/2018/4678362. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

