Insights into the Genomic Regions and Candidate Genes of Senescence-Related Traits in Upland Cotton via GWAS
- PMID: 35955713
- PMCID: PMC9368895
- DOI: 10.3390/ijms23158584
Insights into the Genomic Regions and Candidate Genes of Senescence-Related Traits in Upland Cotton via GWAS
Abstract
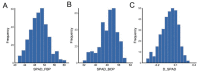
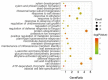
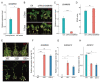
Senescence is the last stage of plant development and is controlled by both internal and external factors. Premature senescence significantly affects the yield and quality of cotton. However, the genetic architecture underlying cotton senescence remains unclear. In this study, genome-wide association studies (GWAS) were performed based on 3,015,002 high-quality SNP markers from the resequencing data of 355 upland cotton accessions to detect genomic regions for cotton senescence. A total of 977 candidate genes within 55 senescence-related genomic regions (SGRs), SGR1-SGR55, were predicted. Gene ontology (GO) analysis of candidate genes revealed that a set of biological processes was enriched, such as salt stress, ethylene processes, and leaf senescence. Furthermore, in the leaf senescence GO term, one candidate gene was focused on: Gohir.A12G270900 (GhMKK9), located in SGR36, which encodes a protein of the MAP kinase kinase family. Quantitative real-time PCR (qRT-PCR) analysis showed that GhMKK9 was up-regulated in old cotton leaves. Overexpression of GhMKK9 in Arabidopsis accelerated natural leaf senescence. Virus-induced gene silencing (VIGS) of GhMKK9 in cotton increased drought tolerance. These results suggest that GhMKK9 is a positive regulator and might be involved in drought-induced senescence in cotton. The results provide new insights into the genetic basis of cotton senescence and will be useful for improving cotton breeding in the future.
Keywords: GWAS; GhMKK9; candidate gene; genomic region; senescence; upland cotton.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Uncovering Novel Genomic Regions and Candidate Genes for Senescence-Related Traits by Genome-Wide Association Studies in Upland Cotton (Gossypium hirsutum L.).Front Plant Sci. 2022 Jan 5;12:809522. doi: 10.3389/fpls.2021.809522. eCollection 2021. Front Plant Sci. 2022. PMID: 35069667 Free PMC article.
-
Analysis of Drought Tolerance and Associated Traits in Upland Cotton at the Seedling Stage.Int J Mol Sci. 2019 Aug 9;20(16):3888. doi: 10.3390/ijms20163888. Int J Mol Sci. 2019. PMID: 31404956 Free PMC article.
-
Genome-wide association study identified genetic variations and candidate genes for plant architecture component traits in Chinese upland cotton.Theor Appl Genet. 2018 Jun;131(6):1299-1314. doi: 10.1007/s00122-018-3079-5. Epub 2018 Mar 1. Theor Appl Genet. 2018. PMID: 29497767
-
Genome-wide association study reveals novel quantitative trait loci and candidate genes of lint percentage in upland cotton based on the CottonSNP80K array.Theor Appl Genet. 2022 Jul;135(7):2279-2295. doi: 10.1007/s00122-022-04111-1. Epub 2022 May 16. Theor Appl Genet. 2022. PMID: 35570221 Review.
-
Recent progression and future perspectives in cotton genomic breeding.J Integr Plant Biol. 2023 Feb;65(2):548-569. doi: 10.1111/jipb.13388. Epub 2022 Dec 31. J Integr Plant Biol. 2023. PMID: 36226594 Review.
Cited by
-
Research on Plant Genomics and Breeding.Int J Mol Sci. 2023 Oct 18;24(20):15298. doi: 10.3390/ijms242015298. Int J Mol Sci. 2023. PMID: 37894978 Free PMC article.
-
Hydrangea arborescens 'Annabelle' Flower Formation and Flowering in the Current Year.Plants (Basel). 2023 Dec 7;12(24):4103. doi: 10.3390/plants12244103. Plants (Basel). 2023. PMID: 38140430 Free PMC article.
-
Research on Plant Genomics and Breeding 2.0.Int J Mol Sci. 2024 Jun 17;25(12):6659. doi: 10.3390/ijms25126659. Int J Mol Sci. 2024. PMID: 38928365 Free PMC article.
-
AmCBF1 activates the expression of GhClpR1 to mediate dark-green leaves in cotton (Gossypium hirsutum).Plant Cell Rep. 2024 Mar 5;43(3):83. doi: 10.1007/s00299-024-03171-5. Plant Cell Rep. 2024. PMID: 38441719
-
Genetic Foundation of Leaf Senescence: Insights from Natural and Cultivated Plant Diversity.Plants (Basel). 2024 Dec 4;13(23):3405. doi: 10.3390/plants13233405. Plants (Basel). 2024. PMID: 39683197 Free PMC article. Review.
References
-
- Shahrajabian M.H., Sun W., Cheng Q. Considering White Gold, Cotton, for its Fiber, Seed Oil, Traditional and Modern Health Benefits. J. Biol. Environ. Sci. 2020;14:25–39.
-
- Gallagher J.P., Grover C.E., Rex K., Moran M., Wendel J.F. A New Species of Cotton from Wake Atoll, Gossypium Stephensii (Malvaceae) Syst. Bot. 2017;42:115–123. doi: 10.1600/036364417X694593. - DOI
-
- Grover C., Zhu X., Grupp K., Jareczek J., Gallagher J., Szadkowski E., Seijo J.G., Wendel J. Molecular Confirmation of Species Status for the Allopolyploid Cotton Species, Gossypium Ekmanianum Wittmack. Genet. Resour. Crop Evol. 2015;62:103–114. doi: 10.1007/s10722-014-0138-x. - DOI
-
- Hulse-Kemp A.M., Lemm J., Plieske J., Ashrafi H., Buyyarapu R., Fang D.D., Frelichowski J., Giband M., Hague S., Hinze L.L., et al. Development of a 63K SNP Array for Cotton and High-Density Mapping of Intraspecific and Interspecific Populations of Gossypium spp. G3 Genes Genomes Genet. 2015;5:1187–1209. doi: 10.1534/g3.115.018416. - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

