Bioinformatics Analysis of the Periodicity in Proteins with Coiled-Coil Structure-Enumerating All Decompositions of Sequence Periods
- PMID: 35955828
- PMCID: PMC9369452
- DOI: 10.3390/ijms23158692
Bioinformatics Analysis of the Periodicity in Proteins with Coiled-Coil Structure-Enumerating All Decompositions of Sequence Periods
Abstract
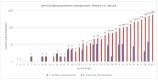
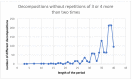
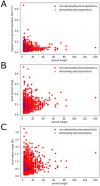
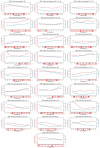
A coiled coil is a structural motif in proteins that consists of at least two α-helices wound around each other. For structural stabilization, these α-helices form interhelical contacts via their amino acid side chains. However, there are restrictions as to the distances along the amino acid sequence at which those contacts occur. As the spatial period of the α-helix is 3.6, the most frequent distances between hydrophobic contacts are 3, 4, and 7. Up to now, the multitude of possible decompositions of α-helices participating in coiled coils at these distances has not been explored systematically. Here, we present an algorithm that computes all non-redundant decompositions of sequence periods of hydrophobic amino acids into distances of 3, 4, and 7. Further, we examine which decompositions can be found in nature by analyzing the available data and taking a closer look at correlations between the properties of the coiled coil and its decomposition. We find that the availability of decompositions allowing for coiled-coil formation without putting too much strain on the α-helix geometry follows an oscillatory pattern in respect of period length. Our algorithm supplies the basis for exploring the possible decompositions of coiled coils of any period length.
Keywords: aldehyde dehydrogenase; coiled coil; decomposition algorithm; hexaprenyl diphosphate synthase; hydrophobic interaction; money-changing problem; recursive algorithm; seduheptulose-7-phosphate isomerase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


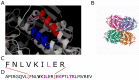

References
-
- Crick F. The packing of α-helices: Simple coiled-coils. Acta Crystallogr. 1953;6:689–697. doi: 10.1107/S0365110X53001964. - DOI
-
- Hodges R.S., Sodek J., Smillie L., Jurasek L. Cold Spring Harbor Symposia on Quantitative Biology. Cold Spring Harbor Laboratory Press; New York, NY, USA: 1979. pp. 299–310. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

