Expression Profiling and MicroRNA Regulatory Networks of Homeobox Family Genes in Sugarcane Saccharum spontaneum L
- PMID: 35955858
- PMCID: PMC9369071
- DOI: 10.3390/ijms23158724
Expression Profiling and MicroRNA Regulatory Networks of Homeobox Family Genes in Sugarcane Saccharum spontaneum L
Abstract
Homeobox (HB) genes play important roles in plant growth and development processes, particularly in the formation of lateral organs. Thus, they could influence leaf morphogenesis and biomass formation in plants. However, little is known about HBs in sugarcane, a crucial sugar crop, due to its complex genetic background. Here, 302 allelic sequences for 104 HBs were identified and divided into 13 subfamilies in sugarcane Saccharum spontaneum. Comparative genomics revealed that whole-genome duplication (WGD)/segmental duplication significantly promoted the expansion of the HB family in S. spontaneum, with SsHB26, SsHB63, SsHB64, SsHB65, SsHB67, SsHB95, and SsHB96 being retained from the evolutionary event before the divergence of dicots and monocots. Based on the analysis of transcriptome and degradome data, we speculated that SsHB15 and SsHB97 might play important roles in regulating sugarcane leaf morphogenesis, with miR166 and SsAGO10 being involved in the regulation of SsHB15 expression. Moreover, subcellular localization and transcriptional activity detection assays demonstrated that these two genes, SsHB15 and SsHB97, were functional transcription factors. This study demonstrated the evolutionary relationship and potential functions of SsHB genes and will enable the further investigation of the functional characterization and the regulatory mechanisms of SsHBs.
Keywords: Saccharum spontaneum; expression analysis; homeobox family; microRNA regulation; phylogenetic analysis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
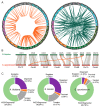






Similar articles
-
Genomic investigation of duplication, functional conservation, and divergence in the LRR-RLK Family of Saccharum.BMC Genomics. 2024 Feb 9;25(1):165. doi: 10.1186/s12864-024-10073-z. BMC Genomics. 2024. PMID: 38336615 Free PMC article.
-
Genome-wide identification and expression profiling of DREB genes in Saccharum spontaneum.BMC Genomics. 2021 Jun 17;22(1):456. doi: 10.1186/s12864-021-07799-5. BMC Genomics. 2021. PMID: 34139993 Free PMC article.
-
Comparative analysis of sucrose phosphate synthase (SPS) gene family between Saccharum officinarum and Saccharum spontaneum.BMC Plant Biol. 2020 Sep 14;20(1):422. doi: 10.1186/s12870-020-02599-7. BMC Plant Biol. 2020. PMID: 32928111 Free PMC article.
-
Comparative genomics revealed the gene evolution and functional divergence of magnesium transporter families in Saccharum.BMC Genomics. 2019 Jan 24;20(1):83. doi: 10.1186/s12864-019-5437-3. BMC Genomics. 2019. PMID: 30678642 Free PMC article.
-
Genome-wide analysis of R2R3-MYB transcription factors family in the autopolyploid Saccharum spontaneum: an exploration of dominance expression and stress response.BMC Genomics. 2021 Aug 18;22(1):622. doi: 10.1186/s12864-021-07689-w. BMC Genomics. 2021. PMID: 34404342 Free PMC article.
Cited by
-
Genomic investigation of duplication, functional conservation, and divergence in the LRR-RLK Family of Saccharum.BMC Genomics. 2024 Feb 9;25(1):165. doi: 10.1186/s12864-024-10073-z. BMC Genomics. 2024. PMID: 38336615 Free PMC article.
-
Enhancement of healthful novel sugar contents in genetically engineered sugarcane juice integrated with molecularly characterized ThSyGII (CEMB-SIG2).Sci Rep. 2022 Nov 3;12(1):18621. doi: 10.1038/s41598-022-23130-y. Sci Rep. 2022. PMID: 36329173 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

