Genomic Insights of First erm B-Positive ST338-SCC mec VT/CC59 Taiwan Clone of Community-Associated Methicillin-Resistant Staphylococcus aureus in Poland
- PMID: 35955887
- PMCID: PMC9369149
- DOI: 10.3390/ijms23158755
Genomic Insights of First erm B-Positive ST338-SCC mec VT/CC59 Taiwan Clone of Community-Associated Methicillin-Resistant Staphylococcus aureus in Poland
Abstract
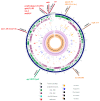
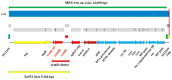
We report the first Polish representative of community-associated methicillin-resistant Staphylococcus aureus (CA-MRSA), lukS/F-PV-positive, encoding the ermB gene, as a genetic determinant of constitutive resistance to macrolides, lincosamides, and streptogramin B antibiotics, cMLS-B. This is the first detection of the CA-MRSA strain responsible for nosocomial infection in the Warsaw Clinical Hospital. Resistance to β-lactams associates with a composite genetic element, SCCmec cassette type VT (5C2&5). We assigned the strain to sequence type ST338 (single-locus variant of ST59), clonal complex CC59, spa-type t437, and agr-type I. Genomic-based comparison was designated SO574/12 as an international Taiwan clone, which has been so far described mainly in the Asia-Pacific region. The ermB gene locates on the chromosome within the 14,690 bp mobile element structure, i.e., the MESPM1-like structure, which also encodes aminoglycoside- and streptothricin-resistance genes. The MESPM1-like structure is a composite transposon containing Tn551, flanked by direct repeats of IS1216V insertion sequences, which probably originates from Enterococcus. The ermB is preceded by the 273 bp regulatory region that contains the regulatory 84 bp ermBL ORF, encoding the 27 amino acid leader peptides. The latest research suggests that a new leader peptide, ermBL2, also exists in the ermB regulatory region. Therefore, the detailed function of ermBL2 requires further investigations.
Keywords: CA-MRSA; IS1216V insertion sequence; Panton–Valentine leukocidin (PVL); ST338/CC59; Taiwan clone; ermB gene; macrolides lincosamides and streptogramin B (MLS-B); mobile element structure (MES PM1).
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Comparative genomics of community-acquired ST59 methicillin-resistant Staphylococcus aureus in Taiwan: novel mobile resistance structures with IS1216V.PLoS One. 2012;7(10):e46987. doi: 10.1371/journal.pone.0046987. Epub 2012 Oct 5. PLoS One. 2012. PMID: 23071689 Free PMC article.
-
Molecular Evolutionary Pathways toward Two Successful Community-Associated but Multidrug-Resistant ST59 Methicillin-Resistant Staphylococcus aureus Lineages in Taiwan: Dynamic Modes of Mobile Genetic Element Salvages.PLoS One. 2016 Sep 8;11(9):e0162526. doi: 10.1371/journal.pone.0162526. eCollection 2016. PLoS One. 2016. PMID: 27606427 Free PMC article.
-
Tracking the evolution of the two successful CC59 methicillin-resistant Staphylococcus aureus clones in Taiwan: the divergence time of the two clades is estimated to be the 1980s.Int J Antimicrob Agents. 2020 Aug;56(2):106047. doi: 10.1016/j.ijantimicag.2020.106047. Epub 2020 Jun 13. Int J Antimicrob Agents. 2020. PMID: 32544568
-
Community-associated meticillin-resistant Staphylococcus aureus in children in Taiwan, 2000s.Int J Antimicrob Agents. 2011 Jul;38(1):2-8. doi: 10.1016/j.ijantimicag.2011.01.011. Epub 2011 Mar 11. Int J Antimicrob Agents. 2011. PMID: 21397461 Review.
-
The evolution of Staphylococcus aureus.Infect Genet Evol. 2008 Dec;8(6):747-63. doi: 10.1016/j.meegid.2008.07.007. Epub 2008 Jul 29. Infect Genet Evol. 2008. PMID: 18718557 Review.
Cited by
-
Comparative De Novo and Pan-Genome Analysis of MDR Nosocomial Bacteria Isolated from Hospitals in Jeddah, Saudi Arabia.Microorganisms. 2023 Sep 28;11(10):2432. doi: 10.3390/microorganisms11102432. Microorganisms. 2023. PMID: 37894090 Free PMC article.
References
-
- Bogut A., Koziol-Montewka M., Baranowicz I., Jozwiak L., Al-Doori Z., Morrison D., Kaczor D., Ksiazek A. Community-acquired methicillin-resistant Staphylococcus aureus (CA-MRSA) in Poland: Further evidence for the changing epidemiology of MRSA. New Microbiol. 2008;31:229–234. - PubMed
-
- Chen J., Luo Y., Zhang S., Liang Z., Wang Y., Zhang Y., Zhou G., Jia Y., Chen L., She D. Community-acquired necrotizing pneumonia caused by methicillin-resistant Staphylococcus aureus producing Panton-Valentine leukocidin in a Chinese teenager: Case report and literature review. Int. J. Infect. Dis. 2014;26:17–21. doi: 10.1016/j.ijid.2014.02.025. - DOI - PubMed
-
- Rolo J., Miragaia M., Turlej-Rogacka A., Empel J., Bouchami O., Faria N.A., Tavares A., Hryniewicz W., Fluit A.C., de Lencastre H., et al. High genetic diversity among community-associated Staphylococcus aureus in Europe: Results from a multicenter study. PLoS ONE. 2012;7:e34768. doi: 10.1371/journal.pone.0034768. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

