Single-cell analysis of transcription factor regulatory networks reveals molecular basis for subtype-specific dysregulation in acute myeloid leukemia
- PMID: 35957668
- PMCID: PMC9362874
- DOI: 10.1097/BS9.0000000000000113
Single-cell analysis of transcription factor regulatory networks reveals molecular basis for subtype-specific dysregulation in acute myeloid leukemia
Abstract
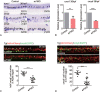
Highly heterogeneous acute myeloid leukemia (AML) exhibits dysregulated transcriptional programs. Transcription factor (TF) regulatory networks underlying AML subtypes have not been elucidated at single-cell resolution. Here, we comprehensively mapped malignancy-related TFs activated in different AML subtypes by analyzing single-cell RNA sequencing data from AMLs and healthy donors. We first identified six modules of regulatory networks which were prevalently dysregulated in all AML patients. AML subtypes featured with different malignant cellular composition possessed subtype-specific regulatory TFs associated with differentiation suppression or immune modulation. At last, we validated that ERF was crucial for the development of hematopoietic stem/progenitor cells by performing loss- and gain-of-function experiments in zebrafish embryos. Collectively, our work thoroughly documents an abnormal spectrum of transcriptional regulatory networks in AML and reveals subtype-specific dysregulation basis, which provides a prospective view to AML pathogenesis and potential targets for both diagnosis and therapy.
Keywords: Acute myeloid leukaemia; Co-expression analysis; Single-cell RNA-sequencing; Transcription factor; Transcriptional regulatory network.
Copyright © 2022 The Authors. Published by Wolters Kluwer Health Inc., on behalf of the Chinese Medical Association (CMA) and Institute of Hematology, Chinese Academy of Medical Sciences & Peking Union Medical College (IHCAMS).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




Similar articles
-
Molecular dysfunctions in acute myeloid leukemia revealed by integrated analysis of microRNA and transcription factor.Int J Oncol. 2016 Jun;48(6):2367-80. doi: 10.3892/ijo.2016.3489. Epub 2016 Apr 15. Int J Oncol. 2016. PMID: 27082628
-
Computational identification of the normal and perturbed genetic networks involved in myeloid differentiation and acute promyelocytic leukemia.Genome Biol. 2008;9(2):R38. doi: 10.1186/gb-2008-9-2-r38. Epub 2008 Feb 21. Genome Biol. 2008. PMID: 18291030 Free PMC article.
-
A single-cell survey of cellular hierarchy in acute myeloid leukemia.J Hematol Oncol. 2020 Sep 25;13(1):128. doi: 10.1186/s13045-020-00941-y. J Hematol Oncol. 2020. PMID: 32977829 Free PMC article.
-
LSD1 inhibition modulates transcription factor networks in myeloid malignancies.Front Oncol. 2023 Mar 10;13:1149754. doi: 10.3389/fonc.2023.1149754. eCollection 2023. Front Oncol. 2023. PMID: 36969082 Free PMC article. Review.
-
Transcriptional networks in acute myeloid leukemia.Genes Chromosomes Cancer. 2019 Dec;58(12):859-874. doi: 10.1002/gcc.22794. Epub 2019 Aug 16. Genes Chromosomes Cancer. 2019. PMID: 31369171 Review.
Cited by
-
Transcription factor genetics and biology in predisposition to bone marrow failure and hematological malignancy.Front Oncol. 2023 Jun 12;13:1183318. doi: 10.3389/fonc.2023.1183318. eCollection 2023. Front Oncol. 2023. PMID: 37377909 Free PMC article. Review.
-
Data-driven modeling of core gene regulatory network underlying leukemogenesis in IDH mutant AML.NPJ Syst Biol Appl. 2024 Apr 9;10(1):38. doi: 10.1038/s41540-024-00366-0. NPJ Syst Biol Appl. 2024. PMID: 38594351 Free PMC article.
-
Classification of acute myeloid leukemia based on multi-omics and prognosis prediction value.Mol Oncol. 2025 Jun;19(6):1836-1854. doi: 10.1002/1878-0261.70000. Epub 2025 Feb 10. Mol Oncol. 2025. PMID: 39927613 Free PMC article.
-
Advances in single-cell RNA sequencing and its applications in cancer research.J Hematol Oncol. 2023 Aug 24;16(1):98. doi: 10.1186/s13045-023-01494-6. J Hematol Oncol. 2023. PMID: 37612741 Free PMC article. Review.
References
-
- Shah A, Andersson TM, Rachet B, Bjorkholm M, Lambert PC. Survival and cure of acute myeloid leukaemia in England, 1971–2006: a population-based study. Br J Haematol 2013;162 (4):509–516. doi:10.1111/bjh.12425. - PubMed
-
- Stavast CJ, Leenen PJM, Erkeland SJ. The interplay between critical transcription factors and microRNAs in the control of normal and malignant myelopoiesis. Cancer Lett 2018;427:28–37. doi:10.1016/j.canlet.2018.04.010. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
