An amateur gut microbial configuration formed in giant panda for striving to digest cellulose in bamboo: Systematic evidence from intestinal digestive enzymes, functional genes and microbial structures
- PMID: 35958139
- PMCID: PMC9363027
- DOI: 10.3389/fmicb.2022.926515
An amateur gut microbial configuration formed in giant panda for striving to digest cellulose in bamboo: Systematic evidence from intestinal digestive enzymes, functional genes and microbial structures
Abstract
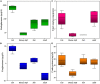
The giant panda has been considered to maximize nutritional intake including protein and soluble carbohydrates in bamboo, but it has spent almost entire life with the high-cellulose diet. Whether giant panda is still helpless about digesting bamboo cellulose or not is always contentious among many researchers around the world. The work has systematically clarified this issue from the perspectives of digestive enzymes, functional genes, and microbial structures in giant panda gut. The intestinal cellulase activities of panda increase with bamboo consumption, performing that the endoglucanase activity of adults reaches 10-fold that of pandas first consuming bamboo. More abundance and types of microbial endoglucanase genes occur in bamboo-diet giant panda gut, and the corresponding GH5 gene cluster is still efficiently transcribed. Gut microbes possessing cellulose-degrading genes, belong to the phylum Firmicutes and some Bacteroidetes, but their structural and functional configurations are insufficient to completely degrade cellulose. Therefore, giant panda is striving to digest cellulose in bamboo, but this adaptation is incomplete. This is probably related to the short straight carnivore-like gut structure of the giant panda, preventing the colonization of some efficient functional but anaerobic-preferred flora.
Keywords: cellulases activity; cellulose digestion; dietary adaptation; giant panda; gut microbiome.
Copyright © 2022 Zhan, Wang, Yao, Zhou, Zhang, Fu, Zhou, Pei and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Succession of Gut Microbial Structure in Twin Giant Pandas During the Dietary Change Stage and Its Role in Polysaccharide Metabolism.Front Microbiol. 2020 Sep 22;11:551038. doi: 10.3389/fmicb.2020.551038. eCollection 2020. Front Microbiol. 2020. PMID: 33072012 Free PMC article.
-
Metagenomic Study Suggests That the Gut Microbiota of the Giant Panda (Ailuropoda melanoleuca) May Not Be Specialized for Fiber Fermentation.Front Microbiol. 2018 Feb 16;9:229. doi: 10.3389/fmicb.2018.00229. eCollection 2018. Front Microbiol. 2018. PMID: 29503636 Free PMC article.
-
The bamboo-eating giant panda harbors a carnivore-like gut microbiota, with excessive seasonal variations.mBio. 2015 May 19;6(3):e00022-15. doi: 10.1128/mBio.00022-15. mBio. 2015. PMID: 25991678 Free PMC article.
-
Progress in Research on the Gut Microflora of the Red Panda (Ailurus fulgens).Microorganisms. 2024 Feb 27;12(3):478. doi: 10.3390/microorganisms12030478. Microorganisms. 2024. PMID: 38543529 Free PMC article. Review.
-
Giant pandas are not an evolutionary cul-de-sac: evidence from multidisciplinary research.Mol Biol Evol. 2015 Jan;32(1):4-12. doi: 10.1093/molbev/msu278. Epub 2014 Oct 1. Mol Biol Evol. 2015. PMID: 25274274 Review.
Cited by
-
Composition, Influencing Factors, and Effects on Host Nutrient Metabolism of Fungi in Gastrointestinal Tract of Monogastric Animals.Animals (Basel). 2025 Mar 1;15(5):710. doi: 10.3390/ani15050710. Animals (Basel). 2025. PMID: 40075993 Free PMC article. Review.
-
Effects of Social Group Housing on the Behavioral and Physiological Responses of Captive Sub-Adult Giant Pandas.Animals (Basel). 2024 Sep 2;14(17):2545. doi: 10.3390/ani14172545. Animals (Basel). 2024. PMID: 39272330 Free PMC article.
References
-
- Alawlaqi M. M., Alharbi A. A. (2020). Exo- and endoglucanase production by Curvularia affinis using bean (Phaseolus vulgaris L.) waste biomass. Bioresour. Bioprocess. 7:6. 10.1186/s40643-020-0296-y - DOI
LinkOut - more resources
Full Text Sources

