Comparative transcriptome analysis unveiling reactive oxygen species scavenging system of Sonneratia caseolaris under salinity stress
- PMID: 35958196
- PMCID: PMC9358527
- DOI: 10.3389/fpls.2022.953450
Comparative transcriptome analysis unveiling reactive oxygen species scavenging system of Sonneratia caseolaris under salinity stress
Abstract
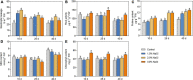
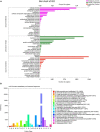
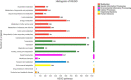
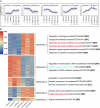
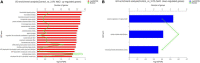
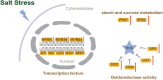
Many mangrove forests have undergone major changes as a result of human activity and global climate change. Sonneratia caseolaris is a common tree located in inner mangroves, and its range extends inland along tidal creeks, as far as the influence of salinity extends. This study investigated the physiological and molecular response mechanisms of S. caseolaris by analyzing its antioxidant defense capacity, including its differentially expressed genes (DEGs) under similar salt stress conditions. Salt treatment significantly affected the osmoprotectants and lipid peroxidation in S. caseolaris seedlings, which increased proline (Pro) content by 31.01-54.90% during all sample periods and decreased malonaldehyde (MDA) content by 12.81 and 18.17% at 25 and 40 days under 3.0% NaCl treatment. Antioxidant enzyme activities increased significantly following 3.0% NaCl treatment. Transcriptome analysis following De novo assembly showed 26,498 matched unigenes. The results showed that 1,263 DEGs responded to transcription factors (TFs) and plant phytohormones and mediated oxidoreductase activity to scavenge reactive oxygen species (ROS) in the control vs. 3.0% NaCl comparison. In addition, the transcription levels of genes associated with auxin and ethylene signal transduction also changed. Under salt stress, ROS scavenging genes (POD, CAT, and APX) and part of AP2, MYB, NAC, C2C2, bHLH, and WRKY TFs were upregulated. This study identified important pathways and candidate genes involved in S. caseolaris salinity tolerance and provided suggestions for further research into the mechanisms of salt tolerance in S. caseolaris.
Keywords: Sonneratia caseolaris; antioxidant; reactive oxygen species; salt stress; transcription factor.
Copyright © 2022 Zhou, Wen, Liao, Lin, Zheng, Li and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
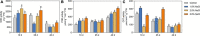

Similar articles
-
Transcriptome analysis of Sonneratia caseolaris seedlings under chilling stress.PeerJ. 2021 Jun 3;9:e11506. doi: 10.7717/peerj.11506. eCollection 2021. PeerJ. 2021. PMID: 34141477 Free PMC article.
-
Comparative transcriptome analyses of a mangrove tree Sonneratia caseolaris and its non-mangrove relatives, Trapa bispinosa and Duabanga grandiflora.Mar Genomics. 2017 Feb;31:13-15. doi: 10.1016/j.margen.2016.10.007. Epub 2016 Oct 31. Mar Genomics. 2017. PMID: 27810366
-
Comparative transcriptome analysis of salt-sensitive and salt-tolerant maize reveals potential mechanisms to enhance salt resistance.Genes Genomics. 2019 Jul;41(7):781-801. doi: 10.1007/s13258-019-00793-y. Epub 2019 Mar 19. Genes Genomics. 2019. PMID: 30887305
-
Understanding salt tolerance mechanism using transcriptome profiling and de novo assembly of wild tomato Solanum chilense.Sci Rep. 2020 Sep 28;10(1):15835. doi: 10.1038/s41598-020-72474-w. Sci Rep. 2020. PMID: 32985535 Free PMC article.
-
Regulation of Reactive Oxygen Species and Antioxidant Defense in Plants under Salinity.Int J Mol Sci. 2021 Aug 28;22(17):9326. doi: 10.3390/ijms22179326. Int J Mol Sci. 2021. PMID: 34502233 Free PMC article. Review.
Cited by
-
De novo transcriptome analysis of high-salinity stress-induced antioxidant activity and plant phytohormone alterations in Sesuvium portulacastrum.Front Plant Sci. 2022 Sep 23;13:995855. doi: 10.3389/fpls.2022.995855. eCollection 2022. Front Plant Sci. 2022. PMID: 36212296 Free PMC article.
-
Functional Characterization of Sugar Beet M14 Antioxidant Enzymes in Plant Salt Stress Tolerance.Antioxidants (Basel). 2022 Dec 27;12(1):57. doi: 10.3390/antiox12010057. Antioxidants (Basel). 2022. PMID: 36670918 Free PMC article.
-
Physiological and molecular response mechanisms of tomato seedlings to cadmium (Cd) and lead (Pb) stress.PeerJ. 2024 Nov 29;12:e18533. doi: 10.7717/peerj.18533. eCollection 2024. PeerJ. 2024. PMID: 39624120 Free PMC article.
-
From swamp to field: how genes from mangroves and its associates can enhance crop salinity tolerance.Mol Biol Rep. 2024 Apr 29;51(1):598. doi: 10.1007/s11033-024-09539-w. Mol Biol Rep. 2024. PMID: 38683409 Review.
References
-
- Agarwal P., Mitra M., Banerjee S., Roy S. (2020). MYB4 transcription factor, a member of R2R3-subfamily of MYB domain protein, regulates cadmium tolerance via enhanced protection against oxidative damage and increases expression of PCS1 and MT1C in Arabidopsis. Plant Sci. 297:110501. 10.1016/j.plantsci.2020.110501 - DOI - PubMed
-
- Biswas P., East A. R., Hewett E. W., Heyes J. A. (2012). Increase in electrolyte leakage as a function of chilling stress and ripening of tomato. Acta Hortic. 945 283–290. 10.17660/ActaHortic.2012.945.37 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

