Functional Annotation of Hypothetical Proteins From the Enterobacter cloacae B13 Strain and Its Association With Pathogenicity
- PMID: 35958299
- PMCID: PMC9358594
- DOI: 10.1177/11779322221115535
Functional Annotation of Hypothetical Proteins From the Enterobacter cloacae B13 Strain and Its Association With Pathogenicity
Abstract
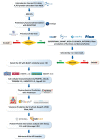
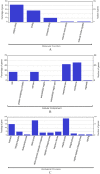
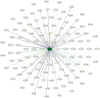
Enterobacter cloacae B13 strain is a rod-shaped gram-negative bacterium that belongs to the Enterobacteriaceae family. It can cause respiratory and urinary tract infections, and is responsible for several outbreaks in hospitals. E. cloacae has become an important pathogen and an emerging global threat because of its opportunistic and multidrug resistant ability. However, little knowledge is present about a large portion of its proteins and functions. Therefore, functional annotation of the hypothetical proteins (HPs) can provide an improved understanding of this organism and its virulence activity. The workflow in the study included several bioinformatic tools which were utilized to characterize functions, family and domains, subcellular localization, physiochemical properties, and protein-protein interactions. The E. cloacae B13 strain has overall 604 HPs, among which 78 were functionally annotated with high confidence. Several proteins were identified as enzymes, regulatory, binding, and transmembrane proteins with essential functions. Furthermore, 23 HPs were predicted to be virulent factors. These virulent proteins are linked to pathogenesis with their contribution to biofilm formation, quorum sensing, 2-component signal transduction or secretion. Better knowledge about the HPs' characteristics and functions will provide a greater overview of the proteome. Moreover, it will help against E. cloacae in neonatal intensive care unit (NICU) outbreaks and nosocomial infections.
Keywords: Enterobacter cloacae; functional annotation; hypothetical proteins; virulence factors.
© The Author(s) 2022.
Conflict of interest statement
Declaration of Conflicting Interests: The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Figures
Similar articles
-
Identification of Novel Abiotic Stress Proteins in Triticum aestivum Through Functional Annotation of Hypothetical Proteins.Interdiscip Sci. 2018 Mar;10(1):205-220. doi: 10.1007/s12539-016-0178-3. Epub 2016 Jul 16. Interdiscip Sci. 2018. PMID: 27421996
-
Functional annotation of hypothetical proteins from the Exiguobacterium antarcticum strain B7 reveals proteins involved in adaptation to extreme environments, including high arsenic resistance.PLoS One. 2018 Jun 25;13(6):e0198965. doi: 10.1371/journal.pone.0198965. eCollection 2018. PLoS One. 2018. PMID: 29940001 Free PMC article.
-
In silico Functional Annotation and Characterization of Hypothetical Proteins from Serratia marcescens FGI94.Biol Bull Russ Acad Sci. 2020;47(4):319-331. doi: 10.1134/S1062359020300019. Epub 2020 Jul 31. Biol Bull Russ Acad Sci. 2020. PMID: 32834707 Free PMC article.
-
The Virulent Hypothetical Proteins: The Potential Drug Target Involved in Bacterial Pathogenesis.Mini Rev Med Chem. 2022;22(20):2608-2623. doi: 10.2174/1389557522666220413102107. Mini Rev Med Chem. 2022. PMID: 35422211 Review.
-
Investigation of an outbreak of Enterobacter cloacae in a neonatal unit and review of the literature.J Hosp Infect. 2008 Sep;70(1):7-14. doi: 10.1016/j.jhin.2008.05.003. Epub 2008 Jul 16. J Hosp Infect. 2008. PMID: 18632183 Review.
Cited by
-
Cross-Over Pathogenic Bacteria Detected in Infected Tomatoes (Solanum lycopersicum L.) and Peppers (Capsicum annuum L.) in Bulgaria.Pathogens. 2022 Dec 9;11(12):1507. doi: 10.3390/pathogens11121507. Pathogens. 2022. PMID: 36558841 Free PMC article.
-
Excavation of Biomarker Candidates for the Diagnosis of Talaromyces marneffei Infection via Genome-Wide Prediction and Functional Annotation of Secreted Proteins.ACS Omega. 2024 Jun 13;9(25):27093-27103. doi: 10.1021/acsomega.4c00571. eCollection 2024 Jun 25. ACS Omega. 2024. PMID: 38947822 Free PMC article.
-
Rapid whole genome sequencing for AMR surveillance in low- and middle-income countries: Oxford Nanopore Technology reveals multidrug-resistant Enterobacter cloacae complex from dairy farms in Sri Lanka.BMC Vet Res. 2025 May 17;21(1):351. doi: 10.1186/s12917-025-04800-1. BMC Vet Res. 2025. PMID: 40382559 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

