Bioinformatics evaluation of a novel angiogenesis related genes-based signature for predicting prognosis and therapeutic efficacy in patients with gastric cancer
- PMID: 35958480
- PMCID: PMC9360876
Bioinformatics evaluation of a novel angiogenesis related genes-based signature for predicting prognosis and therapeutic efficacy in patients with gastric cancer
Abstract
Objective: Tumor angiogenesis plays a pivotal role in the development and metastasis of tumors. This study aimed to elucidate the association between angiogenesis-related genes (ARGs) and the prognosis of patients with gastric cancer (GC).
Methods: Transcriptomics and clinical data of GC samples were obtained from The Cancer Genome Atlas (TCGA) as the training group and those from Gene Expression Omnibus (GEO, including GSE26253, GSE26091 and GSE66229) as the validation groups. Single-sample gene set enrichment analysis (ssGSEA) was performed for gene set enrichment analysis on the gene set of angiogenesis and divided patients into high- or low-ARG group. Subsequently, to improve the availability of the ARG signature, a ARGs subtype predictor was then constructed by integrating of four machine learning methods, including support vector machine (SVM), least absolute shrinkage and selection operator (LASSO) regression, Random Forest and Boruta (RFB) and extreme gradient boosting (XGBoost). Kaplan-Meier and receiver operating characteristic curves were used to evaluate the performance of prognosis prediction. The EPIC and xCELL method were used to calculate the profile of tumor-infiltrated immune cells.
Results: The expression levels of a total of 36 ARGs that correlated with the survival of patients with GC were identified and utilized to establish an ARG-related prognosis signature. The area under the curve for predicting overall survival (OS) in the training group at the 1-, 3- and 5-year was 0.61, 0.64 and 0.76, respectively, and this was further validated using three independent GEO datasets. Moreover, the ARG signatures were significantly correlated with cancer-associated fibroblasts (CAFs), and GC patients that exhibited both high ARG expression level and matrix CAFs level had the most inferior outcomes. The multiple machine learning algorithms were applied to establish a 10-gene ARG subtype predictor, and notably, a high ARG-subtype predictor score was associated with reduced efficacy of immunotherapy, and potential anti-HER2 or FGFR4 therapy, but an increased sensitivity to anti-angiogenesis-related therapy.
Conclusion: The novel ARGs-based classification may act as a potential prognostic predictor for GC and be used as a guidance for clinicians in selecting potential responders for immunotherapy and targeted therapy.
Keywords: Angiogenesis; TCGA; cancer-associated fibroblasts; gastric cancer; immune cell infiltration; immunotherapy.
AJTR Copyright © 2022.
Conflict of interest statement
None.
Figures
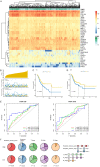






References
-
- Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F. Gastric cancer. Lancet. 2020;396:635–648. - PubMed
-
- Serra O, Galán M, Ginesta MM, Calvo M, Sala N, Salazar R. Comparison and applicability of molecular classifications for gastric cancer. Cancer Treat Rev. 2019;77:29–34. - PubMed
-
- Wang J, Kunzke T, Prade VM, Shen J, Buck A, Feuchtinger A, Haffner I, Luber B, Liu DHW, Langer R, Lordick F, Sun N, Walch A. Spatial metabolomics identifies distinct tumor-specific subtypes in gastric cancer patients. Clin Cancer Res. 2022;21:4383. - PubMed
-
- Carmeliet P. Angiogenesis in life, disease and medicine. Nature. 2005;438:932–936. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
