Hydroxylation Regiochemistry Is Robust to Active Site Mutations in Cytochrome P450cam (CYP101A1)
- PMID: 35960510
- PMCID: PMC9721313
- DOI: 10.1021/acs.biochem.2c00233
Hydroxylation Regiochemistry Is Robust to Active Site Mutations in Cytochrome P450cam (CYP101A1)
Abstract
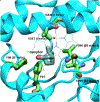
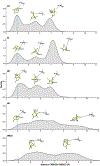
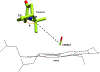
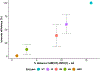
Cytochrome P450cam (CYP101A1) catalyzes the hydroxylation of d-camphor by molecular oxygen. The enzyme-catalyzed hydroxylation exhibits a high degree of regioselectivity and stereoselectivity, with a single major product, d-5-exo-hydroxycamphor, suggesting that the substrate is oriented to facilitate this specificity. In previous work, we used an elastic network model and perturbation response scanning to show that normal deformation modes of the enzyme structure are highly responsive not only to the presence of a substrate but also to the substrate orientation. This work examines the effects of mutations near the active site on substrate localization and orientation. The investigated mutations were designed to promote a change in substrate orientation and/or location that might give rise to different hydroxylation products, while maintaining the same carbon and oxygen atom balances as in the wild type (WT) enzyme. Computational experiments and parallel in vitro site-directed mutations of CYP101A1 were used to examine reaction products and enzyme activity. 1H-15N TROSY-HSQC correlation maps were used to compare the computational results with detectable perturbations in the enzyme structure and dynamics. We found that all of the mutant enzymes retained the same regio- and stereospecificity of hydroxylation as the WT enzyme, with varying degrees of efficiency, which suggests that large portions of the enzyme have been subjected to evolutionary pressure to arrive at the appropriate sequence-structure combination for efficient 5-exo hydroxylation of camphor.
Figures
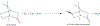

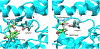



Similar articles
-
Some Surprising Implications of NMR-directed Simulations of Substrate Recognition and Binding by Cytochrome P450cam (CYP101A1).J Mol Biol. 2018 Apr 27;430(9):1295-1310. doi: 10.1016/j.jmb.2018.03.014. Epub 2018 Mar 27. J Mol Biol. 2018. PMID: 29596916 Free PMC article.
-
Hydroxylation of camphor by reduced oxy-cytochrome P450cam: mechanistic implications of EPR and ENDOR studies of catalytic intermediates in native and mutant enzymes.J Am Chem Soc. 2001 Feb 21;123(7):1403-15. doi: 10.1021/ja003583l. J Am Chem Soc. 2001. PMID: 11456714
-
Stereochemistry and deuterium isotope effects in camphor hydroxylation by the cytochrome P450cam monoxygenase system.Biochemistry. 1982 Jan 19;21(2):370-7. doi: 10.1021/bi00531a026. Biochemistry. 1982. PMID: 7074020
-
Differential behavior of the sub-sites of cytochrome 450 active site in binding of substrates, and products (implications for coupling/uncoupling).Biochim Biophys Acta. 2007 Mar;1770(3):360-75. doi: 10.1016/j.bbagen.2006.09.018. Epub 2006 Oct 5. Biochim Biophys Acta. 2007. PMID: 17134838 Review.
-
Updating the Paradigm: Redox Partner Binding and Conformational Dynamics in Cytochromes P450.Acc Chem Res. 2022 Feb 1;55(3):373-380. doi: 10.1021/acs.accounts.1c00632. Epub 2021 Dec 29. Acc Chem Res. 2022. PMID: 34965086 Free PMC article. Review.
Cited by
-
Cooperative Substrate Binding Controls Catalysis in Bacterial Cytochrome P450terp (CYP108A1).J Am Chem Soc. 2023 Feb 13:10.1021/jacs.2c12388. doi: 10.1021/jacs.2c12388. Online ahead of print. J Am Chem Soc. 2023. PMID: 36779970 Free PMC article.
-
Conformational heterogeneity suggests multiple substrate binding modes in CYP106A2.J Inorg Biochem. 2023 Apr;241:112129. doi: 10.1016/j.jinorgbio.2023.112129. Epub 2023 Jan 19. J Inorg Biochem. 2023. PMID: 36731370 Free PMC article.
References
-
- Raag R, and Poulos TL (1989) Crystal structure of the carbon monoxide-substrate-cytochrome P450CAM ternary complex, Biochemistry 28, 7586–7592. - PubMed
-
- Pochapsky SS, Pochapsky TC, and Wei JW (2003) A model for effector activity in a highly specific biological electron transfer complex: The cytochrome P450cam–putidaredoxin couple, Biochemistry 42, 5649–5656. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

