Boost-DiLeu: Enhanced Isobaric N, N-Dimethyl Leucine Tagging Strategy for a Comprehensive Quantitative Glycoproteomic Analysis
- PMID: 35960654
- PMCID: PMC9966376
- DOI: 10.1021/acs.analchem.2c01773
Boost-DiLeu: Enhanced Isobaric N, N-Dimethyl Leucine Tagging Strategy for a Comprehensive Quantitative Glycoproteomic Analysis
Abstract
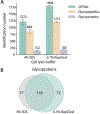
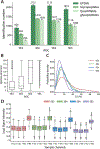
Intact glycopeptide analysis has been of great interest because it can elucidate glycosylation site information and glycan structural composition at the same time. However, mass spectrometry (MS)-based glycoproteomic analysis is hindered by the low abundance and poor ionization efficiency of glycopeptides. Relatively large amounts of starting materials are needed for the enrichment, which makes the identification and quantification of intact glycopeptides from samples with limited quantity more challenging. To overcome these limitations, we developed an improved isobaric labeling strategy with an additional boosting channel to enhance N,N-dimethyl leucine (DiLeu) tagging-based quantitative glycoproteomic analysis, termed as Boost-DiLeu. With the integration of a one-tube sample processing workflow and high-pH fractionation, 3514 quantifiable N-glycopeptides were identified from 30 μg HeLa cell tryptic digests with reliable quantification performance. Furthermore, this strategy was applied to human cerebrospinal fluid (CSF) samples to differentiate N-glycosylation profiles between Alzheimer's disease (AD) patients and non-AD donors. The results revealed processes and pathways affected by dysregulated N-glycosylation in AD, including platelet degranulation, cell adhesion, and extracellular matrix, which highlighted the involvement of N-glycosylation aberrations in AD pathogenesis. Moreover, weighted gene coexpression network analysis (WGCNA) showed nine modules of glycopeptides, two of which were associated with the AD phenotype. Our results demonstrated the feasibility of using this strategy for in-depth glycoproteomic analysis of size-limited clinical samples. Taken together, we developed and optimized a strategy for the enhanced comprehensive quantitative intact glycopeptide analysis with DiLeu labeling, showing significant promise for identifying novel therapeutic targets or biomarkers in biological systems with a limited sample quantity.
Conflict of interest statement
H.Z. has served on scientific advisory boards and/or as a consultant for Abbvie, Alector, Annexon, AZTherapies, CogRx, Denali, Eisai, Nervgen, Pinteon Therapeutics, Red Abbey Labs, Passage Bio, Roche, Samumed, Siemens Healthineers, Triplet Therapeutics, and Wave, has given lectures in symposia sponsored by Cellectricon, Fujirebio, Alzecure and Biogen, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program (outside submitted work). The other authors declare no conflict of interest.
Figures
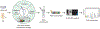

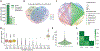

Similar articles
-
Site-specific characterization and quantitation of N-glycopeptides in PKM2 knockout breast cancer cells using DiLeu isobaric tags enabled by electron-transfer/higher-energy collision dissociation (EThcD).Analyst. 2018 May 29;143(11):2508-2519. doi: 10.1039/c8an00216a. Analyst. 2018. PMID: 29687791 Free PMC article.
-
[Recent advances in glycopeptide enrichment and mass spectrometry data interpretation approaches for glycoproteomics analyses].Se Pu. 2021 Oct;39(10):1045-1054. doi: 10.3724/SP.J.1123.2021.06011. Se Pu. 2021. PMID: 34505426 Free PMC article. Review. Chinese.
-
In-depth Site-specific Analysis of N-glycoproteome in Human Cerebrospinal Fluid and Glycosylation Landscape Changes in Alzheimer's Disease.Mol Cell Proteomics. 2021;20:100081. doi: 10.1016/j.mcpro.2021.100081. Epub 2021 Apr 20. Mol Cell Proteomics. 2021. PMID: 33862227 Free PMC article.
-
DiLeu Isobaric Labeling Coupled with Limited Proteolysis Mass Spectrometry for High-Throughput Profiling of Protein Structural Changes in Alzheimer's Disease.Anal Chem. 2023 Jul 4;95(26):9746-9753. doi: 10.1021/acs.analchem.2c05731. Epub 2023 Jun 12. Anal Chem. 2023. PMID: 37307028 Free PMC article.
-
Advances in mass spectrometry-based glycoproteomics: An update covering the period 2017-2021.Electrophoresis. 2022 Jan;43(1-2):370-387. doi: 10.1002/elps.202100188. Epub 2021 Oct 24. Electrophoresis. 2022. PMID: 34614238 Review.
Cited by
-
A Tutorial Review of Labeling Methods in Mass Spectrometry-Based Quantitative Proteomics.ACS Meas Sci Au. 2024 Apr 15;4(4):315-337. doi: 10.1021/acsmeasuresciau.4c00007. eCollection 2024 Aug 21. ACS Meas Sci Au. 2024. PMID: 39184361 Free PMC article. Review.
-
Cotton Ti-IMAC: Developing Phosphorylated Cotton as a Novel Platform for Phosphopeptide Enrichment.ACS Appl Mater Interfaces. 2023 Oct 18;15(41):47893-47901. doi: 10.1021/acsami.3c08697. Epub 2023 Oct 9. ACS Appl Mater Interfaces. 2023. PMID: 37812448 Free PMC article.
-
Quantitative Analysis of Nonhistone Lysine Methylation Sites and Lysine Demethylases in Breast Cancer Cell Lines.J Proteome Res. 2025 Feb 7;24(2):550-561. doi: 10.1021/acs.jproteome.4c00685. Epub 2025 Jan 8. J Proteome Res. 2025. PMID: 39778878
-
Profiling Human Cerebrospinal Fluid (CSF) Endogenous Peptidome in Alzheimer's Disease.Methods Mol Biol. 2024;2758:445-455. doi: 10.1007/978-1-0716-3646-6_24. Methods Mol Biol. 2024. PMID: 38549029
-
Functional Analysis of Mannosyltransferase-Related Genes UvALGs in Ustilaginoidea virens.Int J Mol Sci. 2025 Mar 25;26(7):2979. doi: 10.3390/ijms26072979. Int J Mol Sci. 2025. PMID: 40243604 Free PMC article.
References
-
- Ohtsubo K; Marth JD Glycosylation in Cellular Mechanisms of Health and Disease. Cell 2006, 126, 855–867. - PubMed
-
- Fang P; Xie J; Sang S; Zhang L; Liu M; Yang L; Xu Y; Yan G; Yao J; Gao X; Qian W; Wang Z; Zhang Y; Yang P; Shen H Multilayered N-Glycoproteome Profiling Reveals Highly Heterogeneous and Dysregulated Protein N-Glycosylation Related to Alzheimer’s Disease. Anal. Chem. 2019, 92 (1), 867–874. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

