Differential expression analysis of mRNAs, lncRNAs, and miRNAs expression profiles and construction of ceRNA networks in PEDV infection
- PMID: 35964002
- PMCID: PMC9375197
- DOI: 10.1186/s12864-022-08805-0
Differential expression analysis of mRNAs, lncRNAs, and miRNAs expression profiles and construction of ceRNA networks in PEDV infection
Abstract
Background: Porcine Epidemic Diarrhea Virus (PEDV) is a coronavirus that seriously affects the swine industry. MicroRNAs and long noncoding RNAs are two relevant non-coding RNAs (ncRNAs) class and play crucial roles in a variety of physiological processes. Increased evidence indicates a complex interaction between mRNA and ncRNA. However, our understanding of the function of ncRNA involved in host-PEDV interaction is limited.
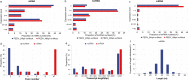
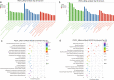
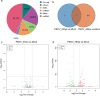
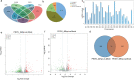
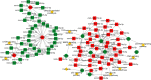
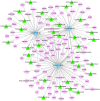
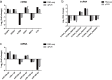
Results: A total of 1,197 mRNA transcripts, 539 lncRNA transcripts, and 208 miRNA transcripts were differentially regulated at 24 h and 48 h post-infection. Gene ontology (GO) and KEGG pathway enrichment analysis showed that DE mRNAs and DE lncRNAs were mainly involved in biosynthesis, innate immunity, and lipid metabolism. Moreover, we constructed a miRNA-mRNA-pathway network using bioinformatics, including 12 DE mRNAs, 120 DE miRNAs, and 11 pathways. Finally, the target genes of DE miRNAs were screened by bioinformatics, and we constructed immune-related lncRNA-miRNA-mRNA ceRNA networks. Then, the selected DE genes were validated by qRT-PCR, which were consistent with the results from RNA-Seq data.
Conclusions: This study provides the comprehensive analysis of the expression profiles of mRNAs, lncRNAs, and miRNAs during PEDV infection. We characterize the ceRNA networks which can provide new insights into the pathogenesis of PEDV.
Keywords: Functional enrichment; PEDV; Signaling pathway; ceRNA network; lncRNA.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

