Identification of TAZ as the essential molecular switch in orchestrating SCLC phenotypic transition and metastasis
- PMID: 35967587
- PMCID: PMC9365451
- DOI: 10.1093/nsr/nwab232
Identification of TAZ as the essential molecular switch in orchestrating SCLC phenotypic transition and metastasis
Erratum in
-
Correction to: Identification of TAZ as the essential molecular switch in orchestrating SCLC phenotypic transition and metastasis.Natl Sci Rev. 2023 Dec 28;10(12):nwac224. doi: 10.1093/nsr/nwac224. eCollection 2023 Dec. Natl Sci Rev. 2023. PMID: 38155833 Free PMC article.
Abstract
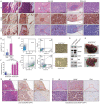
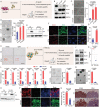
Small-cell lung cancer (SCLC) is a recalcitrant cancer characterized by high metastasis. However, the exact cell type contributing to metastasis remains elusive. Using a Rb1 L/L /Trp53 L/L mouse model, we identify the NCAMhiCD44lo/- subpopulation as the SCLC metastasizing cell (SMC), which is progressively transitioned from the non-metastasizing NCAMloCD44hi cell (non-SMC). Integrative chromatin accessibility and gene expression profiling studies reveal the important role of the SWI/SNF complex, and knockout of its central component, Brg1, significantly inhibits such phenotypic transition and metastasis. Mechanistically, TAZ is silenced by the SWI/SNF complex during SCLC malignant progression, and its knockdown promotes SMC transition and metastasis. Importantly, ectopic TAZ expression reversely drives SMC-to-non-SMC transition and alleviates metastasis. Single-cell RNA-sequencing analyses identify SMC as the dominant subpopulation in human SCLC metastasis, and immunostaining data show a positive correlation between TAZ and patient prognosis. These data uncover high SCLC plasticity and identify TAZ as the key molecular switch in orchestrating SCLC phenotypic transition and metastasis.
Keywords: SWI/SNF complex; TAZ; metastasis; phenotypic transition; small cell lung cancer.
© The Author(s) 2022. Published by Oxford University Press on behalf of China Science Publishing & Media Ltd.
Figures




References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
