Genomic and transcriptomic determinants of response to neoadjuvant therapy in rectal cancer
- PMID: 35970919
- PMCID: PMC9801308
- DOI: 10.1038/s41591-022-01930-z
Genomic and transcriptomic determinants of response to neoadjuvant therapy in rectal cancer
Abstract
The incidence of rectal cancer is increasing in patients younger than 50 years. Locally advanced rectal cancer is still treated with neoadjuvant radiation, chemotherapy and surgery, but recent evidence suggests that patients with a complete response can avoid surgery permanently. To define correlates of response to neoadjuvant therapy, we analyzed genomic and transcriptomic profiles of 738 untreated rectal cancers. APC mutations were less frequent in the lower than in the middle and upper rectum, which could explain the more aggressive behavior of distal tumors. No somatic alterations had significant associations with response to neoadjuvant therapy in a treatment-agnostic manner, but KRAS mutations were associated with faster relapse in patients treated with neoadjuvant chemoradiation followed by consolidative chemotherapy. Overexpression of IGF2 and L1CAM was associated with decreased response to neoadjuvant therapy. RNA-sequencing estimates of immune infiltration identified a subset of microsatellite-stable immune hot tumors with increased response and prolonged disease-free survival.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Figures
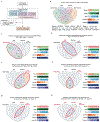
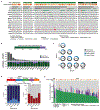








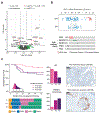

References
-
- Siegel RL, Miller KD, Fuchs HE & Jemal A Cancer Statistics, 2021. CA Cancer J. Clin 71, 7–33 (2021). - PubMed
-
- Garcia-Aguilar J et al. Preliminary results of the organ preservation of rectal adenocarcinoma (OPRA) trial. JCO 38, 4008–4008 (2020).
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

