Gut microbiota individuality is contingent on temporal scale and age in wild meerkats
- PMID: 35975437
- PMCID: PMC9382201
- DOI: 10.1098/rspb.2022.0609
Gut microbiota individuality is contingent on temporal scale and age in wild meerkats
Abstract
Inter-individual differences in gut microbiota composition are hypothesized to generate variation in host fitness-a premise for the evolution of host-gut microbe symbioses. However, recent evidence suggests that gut microbial communities are highly dynamic, challenging the notion that individuals harbour unique gut microbial phenotypes. Leveraging a long-term dataset of wild meerkats, we reconcile these concepts by demonstrating that the relative importance of identity for shaping gut microbiota phenotypes depends on the temporal scale. Across meerkat lifespan, year-to-year variation overshadowed the effects of identity and social group in predicting gut microbiota composition, with identity explaining on average less than 2% of variation. However, identity was the strongest predictor of microbial phenotypes over short sampling intervals (less than two months), predicting on average 20% of variation. The effect of identity was also dependent on meerkat age, with the gut microbiota becoming more individualized and stable as meerkats aged. Nevertheless, while the predictive power of identity was negligible after two months, gut microbiota composition remained weakly individualized compared to that of other meerkats for up to 1 year. These findings illuminate the degree to which individualized gut microbial signatures can be expected, with important implications for the time frames over which gut microbial phenotypes may mediate host physiology, behaviour and fitness in natural populations.
Keywords: gut microbiome; host–microbiota interactions; intraclass correlation coefficient; meerkats; repeatability; temporal dynamics.
Conflict of interest statement
We declare we have no competing interests.
Figures

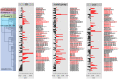

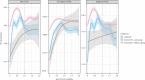
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
