Emergence of Compensatory Mutations Reveals the Importance of Electrostatic Interactions between HIV-1 Integrase and Genomic RNA
- PMID: 35975921
- PMCID: PMC9601147
- DOI: 10.1128/mbio.00431-22
Emergence of Compensatory Mutations Reveals the Importance of Electrostatic Interactions between HIV-1 Integrase and Genomic RNA
Abstract
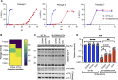
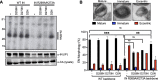
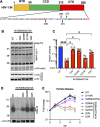
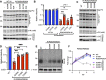
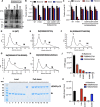
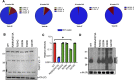
HIV-1 integrase (IN) has a noncatalytic function in virion maturation through its binding to the viral RNA genome (gRNA). Class II IN substitutions inhibit IN-gRNA binding and result in the formation of virions with aberrant morphologies marked by mislocalization of the gRNA between the capsid lattice and the lipid envelope. These viruses are noninfectious due to a block at an early reverse transcription stage in target cells. HIV-1 IN utilizes basic residues within its C-terminal domain (CTD) to bind to the gRNA; however, the molecular nature of how these residues mediate gRNA binding and whether other regions of IN are involved remain unknown. To address this, we have isolated compensatory substitutions in the background of a class II IN mutant virus bearing R269A/K273A substitutions within the IN-CTD. We found that the nearby D256N and D270N compensatory substitutions restored the ability of IN to bind gRNA and led to the formation of mature infectious virions. Reinstating the local positive charge of the IN-CTD through individual D256R, D256K, D278R, and D279R substitutions was sufficient to specifically restore IN-gRNA binding and reverse transcription for the IN R269A/K273A as well as the IN R262A/R263A class II mutants. Structural modeling suggested that compensatory substitutions in the D256 residue created an additional interaction interface for gRNA binding, whereas other substitutions acted locally within the unstructured C-terminal tail of IN. Taken together, our findings highlight the essential role of CTD in gRNA binding and reveal the importance of pliable electrostatic interactions between the IN-CTD and the gRNA. IMPORTANCE In addition to its catalytic function, HIV-1 integrase (IN) binds to the viral RNA genome (gRNA) through positively charged residues (i.e., R262, R263, R269, K273) within its C-terminal domain (CTD) and regulates proper virion maturation. Mutation of these residues results in the formation of morphologically aberrant viruses blocked at an early reverse transcription stage in cells. Here we show that compensatory substitutions in nearby negatively charged aspartic acid residues (i.e., D256N, D270N) restore the ability of IN to bind gRNA for these mutant viruses and result in the formation of accurately matured infectious virions. Similarly, individual charge reversal substitutions at D256 as well as other nearby positions (i.e., D278, D279) are all sufficient to enable the respective IN mutants to bind gRNA, and subsequently restore reverse transcription and virion infectivity. Taken together, our findings reveal the importance of highly pliable electrostatic interactions in IN-gRNA binding.
Keywords: HIV-1; integrase; protein-RNA interactions; virion maturation; virology.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
HIV-1 integrase binding to genomic RNA 5'-UTR induces local structural changes in vitro and in virio.Retrovirology. 2021 Nov 22;18(1):37. doi: 10.1186/s12977-021-00582-0. Retrovirology. 2021. PMID: 34809662 Free PMC article.
-
Capsid Phosphorylation State and Hepadnavirus Virion Secretion.J Virol. 2017 Apr 13;91(9):e00092-17. doi: 10.1128/JVI.00092-17. Print 2017 May 1. J Virol. 2017. PMID: 28228589 Free PMC article.
-
Distribution and Redistribution of HIV-1 Nucleocapsid Protein in Immature, Mature, and Integrase-Inhibited Virions: a Role for Integrase in Maturation.J Virol. 2015 Oct;89(19):9765-80. doi: 10.1128/JVI.01522-15. Epub 2015 Jul 15. J Virol. 2015. PMID: 26178982 Free PMC article.
-
Multimodal Functionalities of HIV-1 Integrase.Viruses. 2022 Apr 28;14(5):926. doi: 10.3390/v14050926. Viruses. 2022. PMID: 35632668 Free PMC article. Review.
-
Going beyond Integration: The Emerging Role of HIV-1 Integrase in Virion Morphogenesis.Viruses. 2020 Sep 9;12(9):1005. doi: 10.3390/v12091005. Viruses. 2020. PMID: 32916894 Free PMC article. Review.
Cited by
-
Defining Distinct RNA-Protein Interactomes of SARS-CoV-2 Genomic and Subgenomic RNAs.J Proteome Res. 2024 Jan 5;23(1):149-160. doi: 10.1021/acs.jproteome.3c00506. Epub 2023 Dec 3. J Proteome Res. 2024. PMID: 38043095 Free PMC article.
-
HIV-1 capsid stability and reverse transcription are finely balanced to minimize sensing of reverse transcription products via the cGAS-STING pathway.mBio. 2024 May 8;15(5):e0034824. doi: 10.1128/mbio.00348-24. Epub 2024 Mar 26. mBio. 2024. PMID: 38530034 Free PMC article.
-
Structural and computational studies of HIV-1 RNA.RNA Biol. 2024 Jan;21(1):1-32. doi: 10.1080/15476286.2023.2289709. Epub 2023 Dec 15. RNA Biol. 2024. PMID: 38100535 Free PMC article. Review.
-
Quinolinonyl Derivatives as Dual Inhibitors of the HIV-1 Integrase Catalytic Site and Integrase-RNA interactions.ACS Med Chem Lett. 2024 Aug 5;15(9):1533-1540. doi: 10.1021/acsmedchemlett.4c00268. eCollection 2024 Sep 12. ACS Med Chem Lett. 2024. PMID: 39291012
-
TAR RNA Mimicry of INI1 and Its Influence on Non-Integration Function of HIV-1 Integrase.Viruses. 2025 May 11;17(5):693. doi: 10.3390/v17050693. Viruses. 2025. PMID: 40431704 Free PMC article. Review.
References
-
- Summa V, Petrocchi A, Bonelli F, Crescenzi B, Donghi M, Ferrara M, Fiore F, Gardelli C, Gonzalez Paz O, Hazuda DJ, Jones P, Kinzel O, Laufer R, Monteagudo E, Muraglia E, Nizi E, Orvieto F, Pace P, Pescatore G, Scarpelli R, Stillmock K, Witmer MV, Rowley M. 2008. Discovery of raltegravir, a potent, selective orally bioavailable HIV-integrase inhibitor for the treatment of HIV-AIDS infection. J Med Chem 51:5843–5855. doi:10.1021/jm800245z. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

