Flexibility of flanking DNA is a key determinant of transcription factor affinity for the core motif
- PMID: 35978548
- PMCID: PMC9674967
- DOI: 10.1016/j.bpj.2022.08.015
Flexibility of flanking DNA is a key determinant of transcription factor affinity for the core motif
Abstract
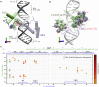
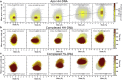
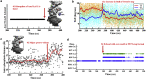
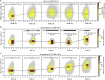
Selective gene regulation is mediated by recognition of specific DNA sequences by transcription factors (TFs). The extremely challenging task of searching out specific cognate DNA binding sites among several million putative sites within the eukaryotic genome is achieved by complex molecular recognition mechanisms. Elements of this recognition code include the core binding sequence, the flanking sequence context, and the shape and conformational flexibility of the composite binding site. To unravel the extent to which DNA flexibility modulates TF binding, in this study, we employed experimentally guided molecular dynamics simulations of ternary complex of closely related Hox heterodimers Exd-Ubx and Exd-Scr with DNA. Results demonstrate that flexibility signatures embedded in the flanking sequences impact TF binding at the cognate binding site. A DNA sequence has intrinsic shape and flexibility features. While shape features are localized, our analyses reveal that flexibility features of the flanking sequences percolate several basepairs and allosterically modulate TF binding at the core. We also show that lack of flexibility in the motif context can render the cognate site resistant to protein-induced shape changes and subsequently lower TF binding affinity. Overall, this study suggests that flexibility-guided DNA shape, and not merely the static shape, is a key unexplored component of the complex DNA-TF recognition code.
Copyright © 2022 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures




Comment in
-
It is in the flanks: Conformational flexibility of transcription factor binding sites.Biophys J. 2022 Oct 18;121(20):3765-3767. doi: 10.1016/j.bpj.2022.09.020. Epub 2022 Sep 21. Biophys J. 2022. PMID: 36182667 Free PMC article. No abstract available.
References
-
- Jones S., van Heyningen P., et al. Thornton J.M. Protein-DNA interactions: a structural analysis. J. Mol. Biol. 1999;287:877–896. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

