Construction of gene modification system with highly efficient and markerless for Monascus ruber M7
- PMID: 35979480
- PMCID: PMC9376451
- DOI: 10.3389/fmicb.2022.952323
Construction of gene modification system with highly efficient and markerless for Monascus ruber M7
Abstract
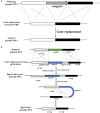
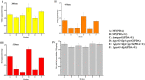
Monascus spp. are traditional medicinal and edible filamentous fungi in China, and can produce various secondary metabolites, such as Monascus pigments (MPs) and citrinin (CIT). Genetic modification methods, such as gene knock-out, complementation, and overexpression, have been used extensively to investigate the function of related genes in Monascus spp.. However, the resistance selection genes that can have been used for genetic modification in Monascus spp. are limited, and the gene replacement frequency (GRF) is usually <5%. Therefore, we are committed to construct a highly efficient gene editing system without resistance selection marker gene. In this study, using M. ruber M7 as the starting strain, we successfully constructed a so-called markerlessly and highly genetic modification system including the mutants ΔmrpyrGΔmrlig4 and ΔmrpyrGΔmrlig4::mrpyrG, in which we used the endogenous gene mrpyrG from M. ruber M7 instead of the resistance marker gene as the screening marker, and simultaneously deleted mrlig4 related to non-homologous end joining in M. ruber M7. Then, the morphology, the growth rate, the production of MPs and CIT of the mutants were analyzed. And the results show that the mutant strains have normal mycelia, cleistothecia and conidia on PDA+Uridine(U) plate, the biomass of each mutant is also no different from M. ruber M7. However, the U addition also has a certain effect on the orange and red pigments yield of M. ruber M7, which needs our further study. Finally, we applied the system to delete multiple genes from M. ruber M7 separately or continuously without any resistance marker gene, and found that the average GRF of ΔmrpyrGΔmrlig4 was about 18 times of that of M. ruber M7. The markerlessly and highly genetic modification system constructed in current study not only will be used for multi-gene simultaneous modification in Monascus spp., and also lays a foundation for investigating the effects of multi-genes modification on Monascus spp..
Keywords: Monascus ruber M7; genetic modification system; mrlig4; mrpyrG; resistance selection marker.
Copyright © 2022 Xu, Li and Chen.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
LinkOut - more resources
Full Text Sources
Miscellaneous

